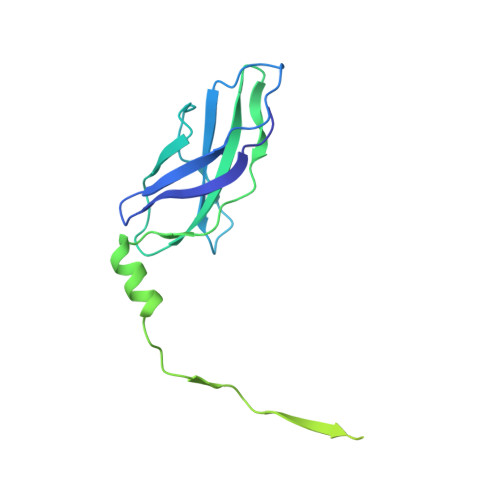
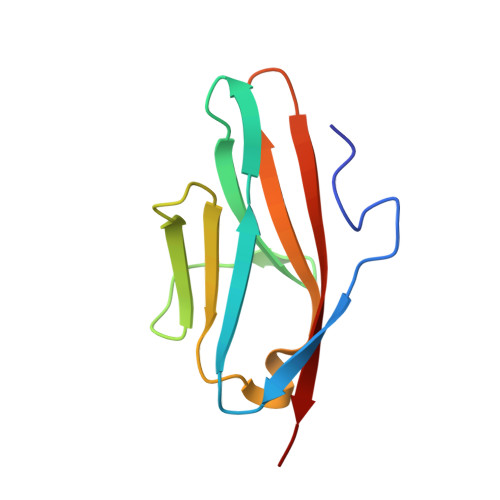
Binding of Myomesin to Obscurin-Like-1 at the Muscle M-Band Provides a Strategy for Isoform-Specific Mechanical Protection.
Pernigo, S., Fukuzawa, A., Beedle, A.E., Holt, M., Round, A., Pandini, A., Garcia-Manyes, S., Gautel, M., Steiner, R.A.(2017) Structure 25: 107-120
- PubMed: 27989621 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2016.11.015
- Primary Citation Related Structures:
5FM4, 5FM5, 5FM8 - PubMed Abstract:
The sarcomeric cytoskeleton is a network of modular proteins that integrate mechanical and signaling roles. Obscurin, or its homolog obscurin-like-1, bridges the giant ruler titin and the myosin crosslinker myomesin at the M-band. Yet, the molecular mechanisms underlying the physical obscurin(-like-1):myomesin connection, important for mechanical integrity of the M-band, remained elusive. Here, using a combination of structural, cellular, and single-molecule force spectroscopy techniques, we decode the architectural and functional determinants defining the obscurin(-like-1):myomesin complex. The crystal structure reveals a trans-complementation mechanism whereby an incomplete immunoglobulin-like domain assimilates an isoform-specific myomesin interdomain sequence. Crucially, this unconventional architecture provides mechanical stability up to forces of ∼135 pN. A cellular competition assay in neonatal rat cardiomyocytes validates the complex and provides the rationale for the isoform specificity of the interaction. Altogether, our results reveal a novel binding strategy in sarcomere assembly, which might have implications on muscle nanomechanics and overall M-band organization.
- Randall Division of Cell and Molecular Biophysics, King's College London, London SE1 1UL, UK.
Organizational Affiliation: