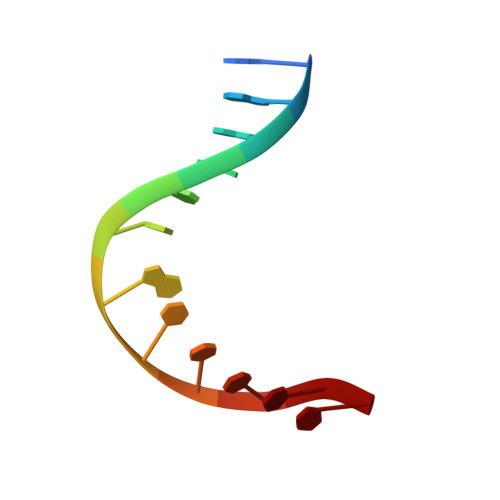
Metal Ion Binding at the Catalytic Site Induces Widely Distributed Changes in a Sequence Specific Protein-DNA Complex.
Sinha, K., Sangani, S.S., Kehr, A.D., Rule, G.S., Jen-Jacobson, L.(2016) Biochemistry 55: 6115-6132
- PubMed: 27786446 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.6b00919
- Primary Citation Related Structures:
5F8A, 5HLK - PubMed Abstract:
Metal ion cofactors can alter the energetics and specificity of sequence specific protein-DNA interactions, but it is unknown if the underlying effects on structure and dynamics are local or dispersed throughout the protein-DNA complex. This work uses EcoRV endonuclease as a model, and catalytically inactive lanthanide ions, which replace the Mg 2+ cofactor. Nuclear magnetic resonance (NMR) titrations indicate that four Lu 3+ or two La 3+ cations bind, and two new crystal structures confirm that Lu 3+ binding is confined to the active sites. NMR spectra show that the metal-free EcoRV complex with cognate (GATATC) DNA is structurally distinct from the nonspecific complex, and that metal ion binding sites are not assembled in the nonspecific complex. NMR chemical shift perturbations were determined for 1 H- 15 N amide resonances, for 1 H- 13 C Ile-δ-CH 3 resonances, and for stereospecifically assigned Leu-δ-CH 3 and Val-γ-CH 3 resonances. Many chemical shifts throughout the cognate complex are unperturbed, so metal binding does not induce major conformational changes. However, some large perturbations of amide and side chain methyl resonances occur as far as 34 Å from the metal ions. Concerted changes in specific residues imply that local effects of metal binding are propagated via a β-sheet and an α-helix. Both amide and methyl resonance perturbations indicate changes in the interface between subunits of the EcoRV homodimer. Bound metal ions also affect amide hydrogen exchange rates for distant residues, including a distant subdomain that contacts DNA phosphates and promotes DNA bending, showing that metal ions in the active sites, which relieve electrostatic repulsion between protein and DNA, cause changes in slow dynamics throughout the complex.
- Department of Biological Sciences, University of Pittsburgh , Pittsburgh, Pennsylvania 15260, United States.
Organizational Affiliation: