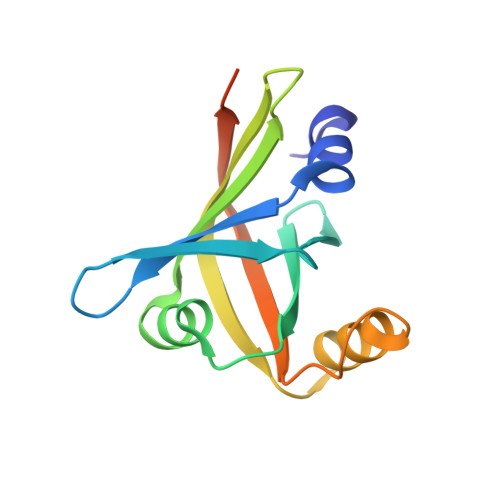
In vitro heme biotransformation by the HupZ enzyme from Group A streptococcus.
Sachla, A.J., Ouattara, M., Romero, E., Agniswamy, J., Weber, I.T., Gadda, G., Eichenbaum, Z.(2016) Biometals 29: 593-609
- PubMed: 27154580 Search on PubMed
- DOI: https://doi.org/10.1007/s10534-016-9937-1
- Primary Citation Related Structures:
5ESC - PubMed Abstract:
In Group A streptococcus (GAS), the metallorepressor MtsR regulates iron homeostasis. Here we describe a new MtsR-repressed gene, which we named hupZ (heme utilization protein). A recombinant HupZ protein was purified bound to heme from Escherichia coli grown in the presence of 5-aminolevulinic acid and iron. HupZ specifically binds heme with stoichiometry of 1:1. The addition of NADPH to heme-bound HupZ (in the presence of cytochrome P450 reductase, NADPH-regeneration system and catalase) triggered progressive decrease of the HupZ Soret band and the appearance of an absorption peak at 660 nm that was resistance to hydrolytic conditions. No spectral changes were observed when ferredoxin and ferredoxin reductase were used as redox partners. Differential spectroscopy with myoglobin or with the ferrous chelator, ferrozine, confirmed that carbon monoxide and free iron are produced during the reaction. ApoHupZ was crystallized as a homodimer with a split β-barrel conformation in each monomer comprising six β strands and three α helices. This structure resembles the split β-barrel domain shared by the members of a recently described family of heme degrading enzymes. However, HupZ is smaller and lacks key residues found in the proteins of the latter group. Phylogenetic analysis places HupZ on a clade separated from those for previously described heme oxygenases. In summary, we have identified a new GAS enzyme-containing split β-barrel and capable of heme biotransformation in vitro; to the best of our knowledge, this is the first enzyme among Streptococcus species with such activity.
- Department of Biology, College of Arts and Sciences, Georgia State University, P.O. Box 4010, Atlanta, GA, 30302-4010, USA.
Organizational Affiliation: