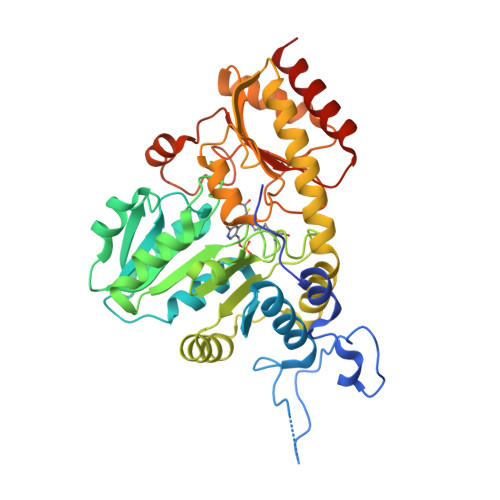
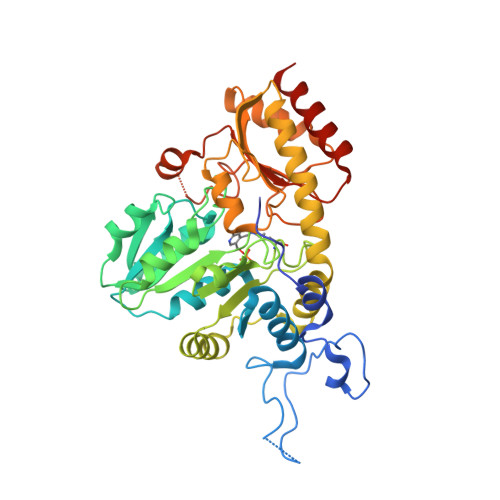
Systemic Depletion of Serum L-Cyst(e)ine with an Engineered Human Enzyme Mediates Potent Induction of ROS and Cancer Ablation
Cramer, S., Saha, A., Tadi, S., Tiziani, S., Yan, W., Triplett, K., Lamb, C., Alters, S., Johnson, D., Zhang, Y., DiGiovanni, J., Georgiou, G., Stone, E.To be published.