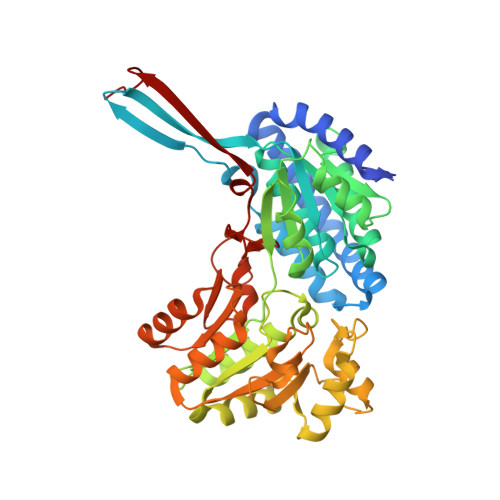
Insight into Coenzyme A cofactor binding and the mechanism of acyl-transfer in an acylating aldehyde dehydrogenase from Clostridium phytofermentans.
Tuck, L.R., Altenbach, K., Ang, T.F., Crawshaw, A.D., Campopiano, D.J., Clarke, D.J., Marles-Wright, J.(2016) Sci Rep 6: 22108-22108
- PubMed: 26899032 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep22108
- Primary Citation Related Structures:
4C3S, 5DBV, 5DRU - PubMed Abstract:
The breakdown of fucose and rhamnose released from plant cell walls by the cellulolytic soil bacterium Clostridium phytofermentans produces toxic aldehyde intermediates. To enable growth on these carbon sources, the pathway for the breakdown of fucose and rhamnose is encapsulated within a bacterial microcompartment (BMC). These proteinaceous organelles sequester the toxic aldehyde intermediates and allow the efficient action of acylating aldehyde dehydrogenase enzymes to produce an acyl-CoA that is ultimately used in substrate-level phosphorylation to produce ATP. Here we analyse the kinetics of the aldehyde dehydrogenase enzyme from the fucose/rhamnose utilisation BMC with different short-chain fatty aldehydes and show that it has activity against substrates with up to six carbon atoms, with optimal activity against propionaldehyde. We have also determined the X-ray crystal structure of this enzyme in complex with CoA and show that the adenine nucleotide of this cofactor is bound in a distinct pocket to the same group in NAD(+). This work is the first report of the structure of CoA bound to an aldehyde dehydrogenase enzyme and our crystallographic model provides important insight into the differences within the active site that distinguish the acylating from non-acylating aldehyde dehydrogenase enzymes.
- Institute of Quantitative Biology, Biochemistry and Biotechnology, The University of Edinburgh, Max Born Crescent, EH9 3BF, UK.
Organizational Affiliation: