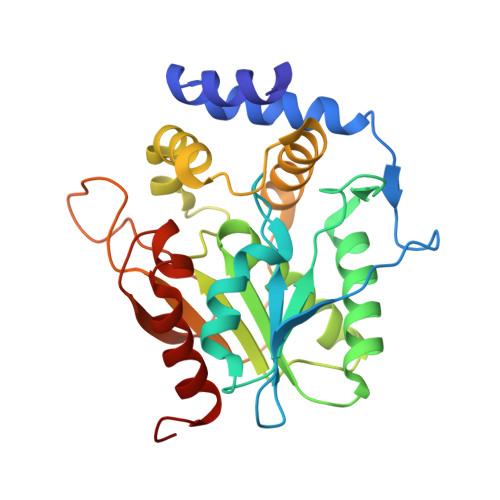
Towards a characterization of the structural determinants of specificity in the macrocyclizing thioesterase for deoxyerythronolide B biosynthesis.
Argyropoulos, P., Bergeret, F., Pardin, C., Reimer, J.M., Pinto, A., Boddy, C.N., Schmeing, T.M.(2015) Biochim Biophys Acta 1860: 486-497
- PubMed: 26592346
- DOI: https://doi.org/10.1016/j.bbagen.2015.11.007
- Primary Citation Related Structures:
5D3K, 5D3Z - PubMed Abstract:
Type I polyketide synthases (PKSs) are giant multidomain proteins that synthesize many therapeutics and other natural products. The synthesis proceeds by a thiotemplate mechanism whereby intermediates are covalently attached to the PKS. The release of the final polyketide is catalyzed by the terminal thioesterase (TE) domain through hydrolysis, transesterification, or macrocyclization. The PKS 6-deoxyerythronolide B synthase (DEBS) produces the 14-membered macrolide core of the clinically important antibiotic erythromycin. The TE domain of DEBS (DEBS TE) has well-established, empirically-defined specificities for hydrolysis or macrocyclization of native and modified substrates. We present efforts towards understanding the structural basis for the specificity of the thioesterase reaction in DEBS TE using a set of novel diphenyl alkylphosphonates, which mimic substrates that are specifically cyclized or hydrolyzed by DEBS TE. We have determined structures of a new construct of DEBS TE alone at 1.7Å, and DEBS TE bound with a simple allylphosphonate at 2.1Å resolution. Other, more complex diphenyl alkylphosphonates inhibit DEBS TE, but we were unable to visualize these faithful cyclization analogs in complex with DEBS TE. This work represents a first step towards using DEBS TE complexed with sophisticated substrate analogs to decipher the specificity determinants in this important reaction.
- Department of Chemistry and Biomolecular Sciences, Centre for Catalysis Research and Innovation, University of Ottawa, Ottawa, ON, K1N 6N5, Canada.
Organizational Affiliation: