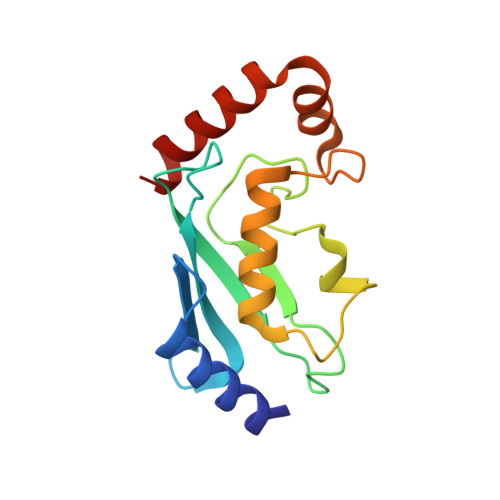
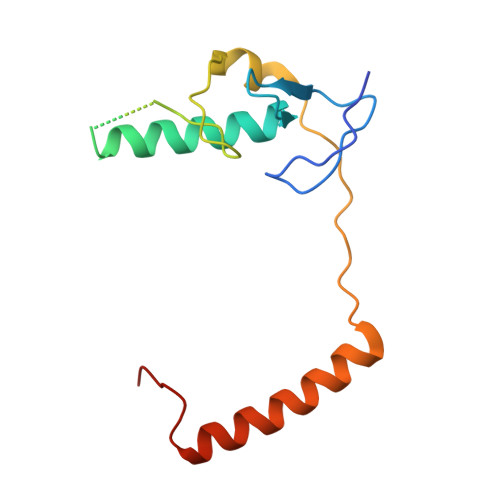
Insights into Ubiquitination from the Unique Clamp-like Binding of the RING E3 AO7 to the E2 UbcH5B.
Li, S., Liang, Y.H., Mariano, J., Metzger, M.B., Stringer, D.K., Hristova, V.A., Li, J., Randazzo, P.A., Tsai, Y.C., Ji, X., Weissman, A.M.(2015) J Biological Chem 290: 30225-30239
- PubMed: 26475854 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M115.685867
- Primary Citation Related Structures:
5D1K, 5D1L, 5D1M - PubMed Abstract:
RING proteins constitute the largest class of E3 ubiquitin ligases. Unlike most RINGs, AO7 (RNF25) binds the E2 ubiquitin-conjugating enzyme, UbcH5B (UBE2D2), with strikingly high affinity. We have defined, by co-crystallization, the distinctive means by which AO7 binds UbcH5B. AO7 contains a structurally unique UbcH5B binding region (U5BR) that is connected by an 11-amino acid linker to its RING domain, forming a clamp surrounding the E2. The U5BR interacts extensively with a region of UbcH5B that is distinct from both the active site and the RING-interacting region, referred to as the backside of the E2. An apparent paradox is that the high-affinity binding of the AO7 clamp to UbcH5B, which is dependent on the U5BR, decreases the rate of ubiquitination. We establish that this is a consequence of blocking the stimulatory, non-covalent, binding of ubiquitin to the backside of UbcH5B. Interestingly, when non-covalent backside ubiquitin binding cannot occur, the AO7 clamp now enhances the rate of ubiquitination. The high-affinity binding of the AO7 clamp to UbcH5B has also allowed for the co-crystallization of previously described and functionally important RING mutants at the RING-E2 interface. We show that mutations having marked effects on function only minimally affect the intermolecular interactions between the AO7 RING and UbcH5B, establishing a high degree of complexity in activation through the RING-E2 interface.
- From the Laboratory of Protein Dynamics and Signaling.
Organizational Affiliation: