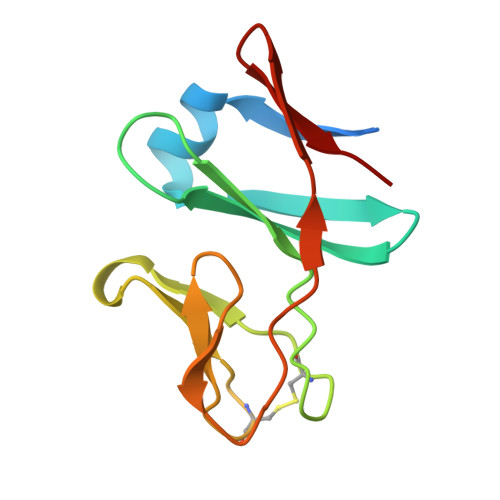
Structural and functional characterisation of the cyanobacterial PetC3 Rieske protein family.
Veit, S., Takeda, K., Tsunoyama, Y., Baymann, F., Nevo, R., Reich, Z., Rogner, M., Miki, K., Rexroth, S.(2016) Biochim Biophys Acta 1857: 1879-1891
- PubMed: 27663073 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbabio.2016.09.007
- Primary Citation Related Structures:
5CXM - PubMed Abstract:
The cyanobacterium Synechocystis PCC 6803 possesses three Rieske isoforms: PetC1, PetC2 and PetC3. While PetC1 and PetC2 have been identified as alternative subunits of the cytochrome b 6 f complex (b 6 f), PetC3 was localized exclusively within the plasma membrane. The spatial separation of PetC3 from the photosynthetic and respiratory protein complexes raises doubt in its involvement in bioenergetic electron transfer. Here we report a detailed structural and functional characterization of the cyanobacterial PetC3 protein family indicating that PetC3 is not a component of the b 6 f and the photosynthetic electron transport as implied by gene annotation. Instead PetC3 has a distinct function in cell envelope homeostasis. Especially proteomic analysis shows that deletion of petC3 in Synechocystis PCC 6803 primarily affects cell envelope proteins including many nutrient transport systems. Therefore, the observed downregulation in the photosynthetic electron transport - mainly caused by photosystem 2 inactivation - might constitute a stress adaptation. Comprehensive in silico sequence analyses revealed that PetC3 proteins are periplasmic lipoproteins tethered to the plasma membrane with a subclass consisting of soluble periplasmic proteins, i.e. their N-terminal domain is inconsistent with their integration into the b 6 f. For the first time, the structure of PetC3 was determined by X-ray crystallography at an atomic resolution revealing significant high similarities to non-b 6 f Rieske subunits in contrast to PetC1. These results suggest that PetC3 affects processes in the periplasmic compartment that only indirectly influence photosynthetic electron transport. For this reason, we suggest to rename "Photosynthetic electron transport Chain 3" (PetC3) proteins as "periplasmic Rieske proteins" (Prp).
- Plant Biochemistry, Faculty of Biology & Biotechnology, Ruhr University of Bochum, 44780 Bochum, Germany.
Organizational Affiliation: