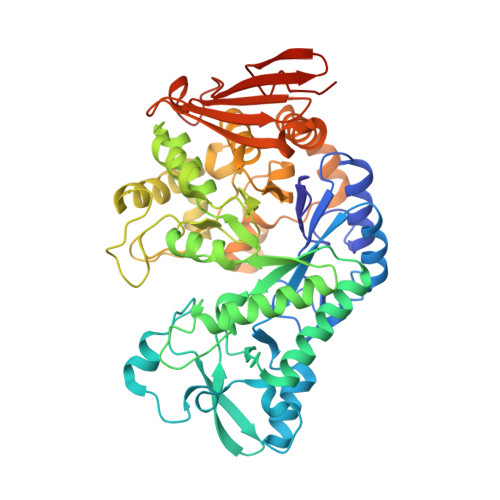
Redesign of the Active Site of Sucrose Phosphorylase through a Clash-Induced Cascade of Loop Shifts.
Kraus, M., Grimm, C., Seibel, J.(2016) Chembiochem 17: 33-36
- PubMed: 26527586
- DOI: https://doi.org/10.1002/cbic.201500514
- Primary Citation of Related Structures:
5C8B - PubMed Abstract:
Sucrose phosphorylases have been applied in the enzymatic production of glycosylated compounds for decades. However, several desirable acceptors, such as flavonoids or stilbenoids, that exhibit diverse antimicrobial, anticarcinogenic or antioxidant properties, remain poor substrates. The Q345F exchange in sucrose phosphorylase from Bifidobacterium adolescentis allows efficient glucosylation of resveratrol, (+)-catechin and (-)-epicatechin in yields of up to 97 % whereas the wild-type enzyme favours sucrose hydrolysis. Three previously undescribed products are made available. The crystal structure of the variant reveals a widened access channel with a hydrophobic aromatic surface that is likely to contribute to the improved activity towards aromatic acceptors. The generation of this channel can be explained in terms of a cascade of structural changes arising from the Q345F exchange. The observed mechanisms are likely to be relevant for the design of other tailor-made enzymes.
- Department of Organic Chemistry, University of Würzburg, Am Hubland, 97074, Würzburg, Germany.
Organizational Affiliation: