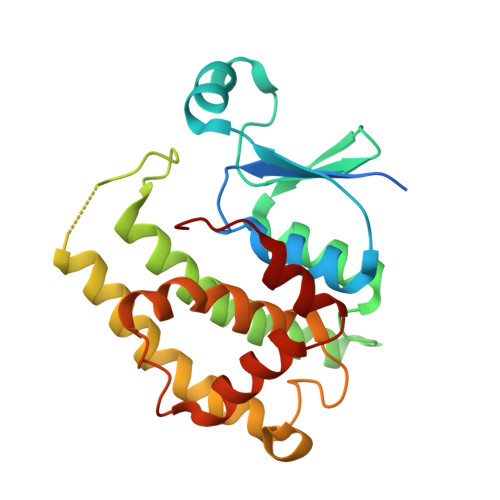
Structure of a Highly Active Cephalopod S-crystallin Mutant: New Molecular Evidence for Evolution from an Active Enzyme into Lens-Refractive Protein.
Tan, W.H., Cheng, S.C., Liu, Y.T., Wu, C.G., Lin, M.H., Chen, C.C., Lin, C.H., Chou, C.Y.(2016) Sci Rep 6: 31176-31176
- PubMed: 27499004
- DOI: https://doi.org/10.1038/srep31176
- Primary Citation of Related Structures:
5B7C - PubMed Abstract:
Crystallins are found widely in animal lenses and have important functions due to their refractive properties. In the coleoid cephalopods, a lens with a graded refractive index provides good vision and is required for survival. Cephalopod S-crystallin is thought to have evolved from glutathione S-transferase (GST) with various homologs differentially expressed in the lens. However, there is no direct structural information that helps to delineate the mechanisms by which S-crystallin could have evolved. Here we report the structural and biochemical characterization of novel S-crystallin-glutathione complex. The 2.35-Å crystal structure of a S-crystallin mutant from Octopus vulgaris reveals an active-site architecture that is different from that of GST. S-crystallin has a preference for glutathione binding, although almost lost its GST enzymatic activity. We've also identified four historical mutations that are able to produce a "GST-like" S-crystallin that has regained activity. This protein recapitulates the evolution of S-crystallin from GST. Protein stability studies suggest that S-crystallin is stabilized by glutathione binding to prevent its aggregation; this contrasts with GST-σ, which do not possess this protection. We suggest that a tradeoff between enzyme activity and the stability of the lens protein might have been one of the major driving force behind lens evolution.
- Department of Life Sciences and Institute of Genome Sciences, National Yang-Ming University, Taipei 112, Taiwan.
Organizational Affiliation: