Turnover of Bacterial Cell Wall by Sltb3, a Multidomain Lytic Transglycosylase of Pseudomonas Aeruginosa.
Lee, M., Dominguez-Gil, T., Hesek, D., Mahasenan, K.V., Lastochkin, E., Hermoso, J.A., Mobashery, S.(2016) ACS Chem Biol 11: 1525
- PubMed: 27035839
- DOI: https://doi.org/10.1021/acschembio.6b00194
- Primary Citation Related Structures:
5ANZ, 5AO7, 5AO8 - PubMed Abstract:
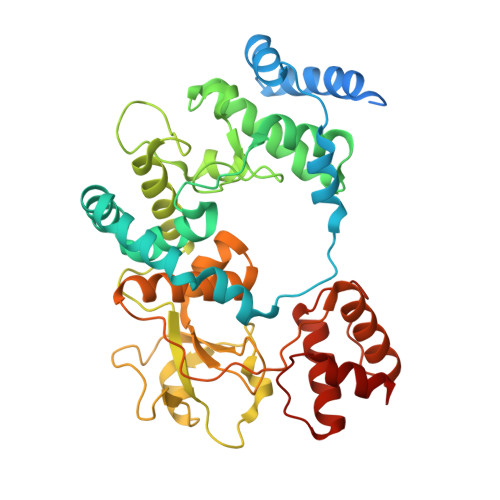
A family of 11 lytic transglycosylases in Pseudomonas aeruginosa, an opportunistic human pathogen, turn over the polymeric bacterial cell wall in the course of its recycling, repair, and maturation. The functions of these enzymes are not fully understood. We disclose herein that SltB3 of P. aeruginosa is an exolytic lytic transglycosylase. We characterize its reaction and its products by the use of peptidoglycan-based molecules. The enzyme recognizes a minimum of four sugars in its substrate but can process a substrate comprised of a peptidoglycan of 20 sugars. The ultimate product of the reaction is N-acetylglucosamine-1,6-anhydro-N-acetylmuramic acid. The X-ray structure of this enzyme is reported for the first time. The enzyme is comprised of four domains, arranged within an annular conformation. The polymeric linear peptidoglycan substrate threads through the opening of the annulus, as it experiences turnover.
- Department of Chemistry and Biochemistry, University of Notre Dame , Nieuwland Science Hall, Notre Dame, Indiana 46556, United States.
Organizational Affiliation: