An Unusual Natural Product Primary Sulfonamide: Synthesis, Carbonic Anhydrase Inhibition and Protein X-Ray Structures of Psammaplin C.
Mujumdar, P., Teruya, K., Tonissen, K.F., Vullo, D., Supuran, C.T., Peat, T.S., Poulsen, S.(2016) J Med Chem 59: 5462
- PubMed: 27172398 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.6b00443
- Primary Citation Related Structures:
5A6H, 5G01, 5G03, 5G0B, 5G0C - PubMed Abstract:
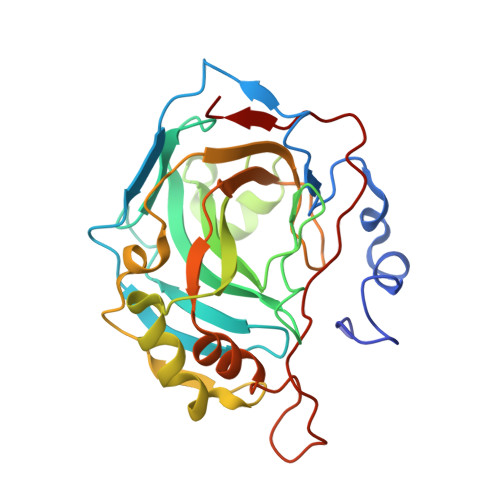
Psammaplin C is one of only two described natural product primary sulfonamides. Here we report the synthesis of psammaplin C and evaluate the inhibition profile against therapeutically relevant carbonic anhydrase (CA) zinc metalloenzymes. The compound exhibited unprecedented inhibition of an important cancer-associated isozyme, hCA XII, with a Ki of 0.79 nM. The compound also displayed good isoform selectivity for hCA XII over other CAs. We present the first reported protein X-ray crystal structures of psammaplin C in complex with human CAs. We engineered the easily crystallized hCA II enzyme to mimic both the hCA IX and hCA XII binding sites and then utilized protein X-ray crystallography to determine the binding pose of psammaplin C within the hCA II, hCA IX, and hCA XII mimic active sites, all to high resolution. This is the first time a natural product primary sulfonamide inhibitor has been assessed for inhibition and binding to CAs.
- Eskitis Institute for Drug Discovery, Griffith University , Don Young Road, Nathan, Queensland 4111, Australia.
Organizational Affiliation: