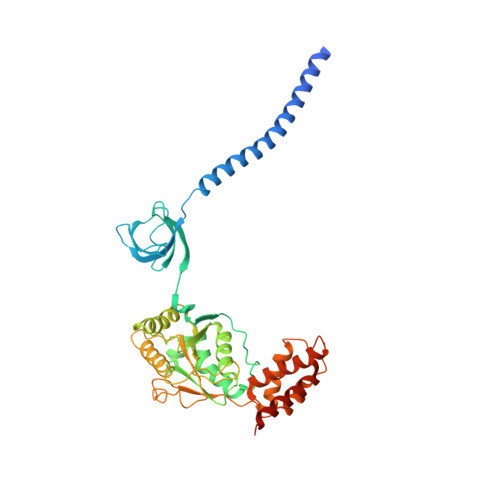
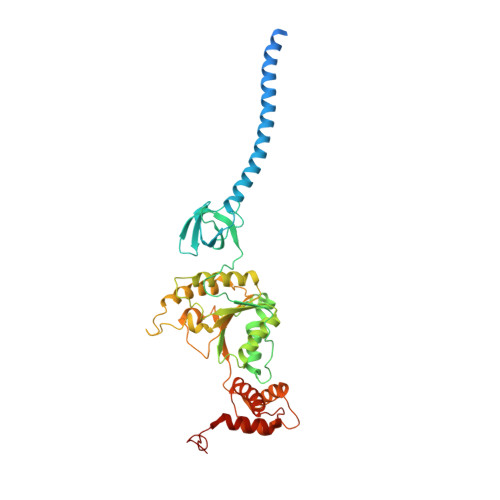
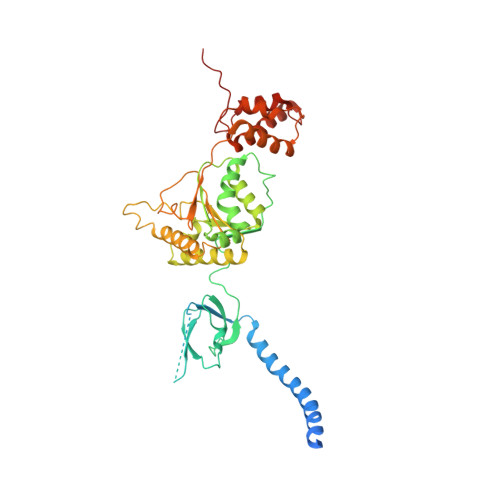
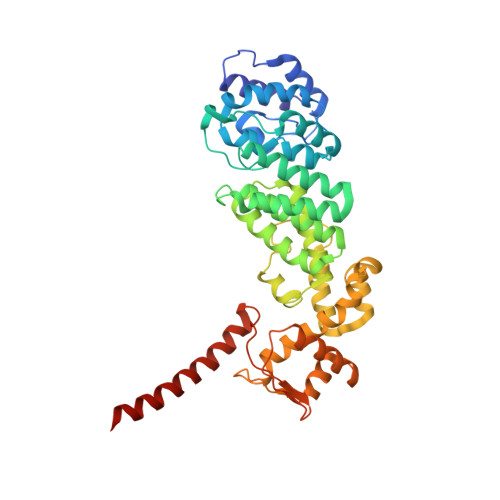
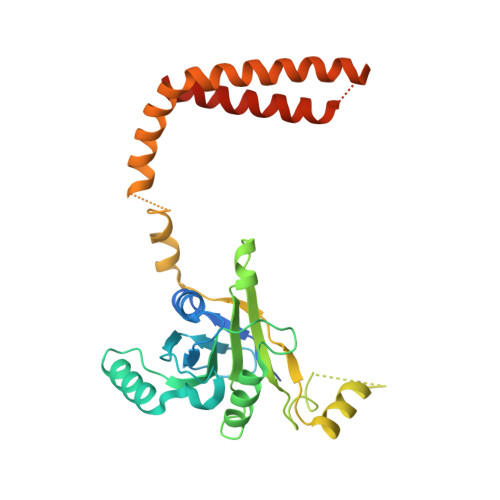
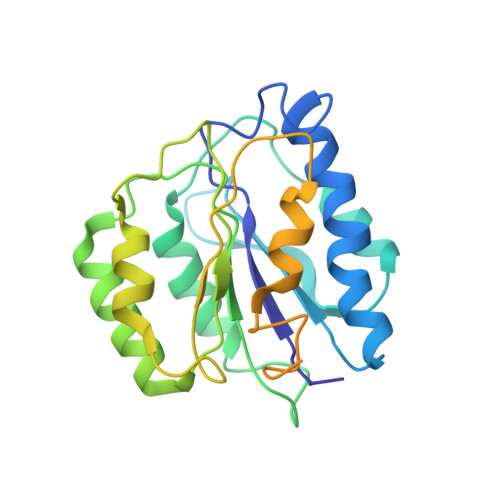
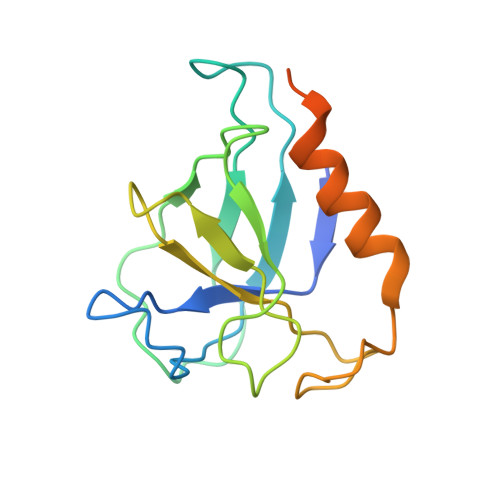
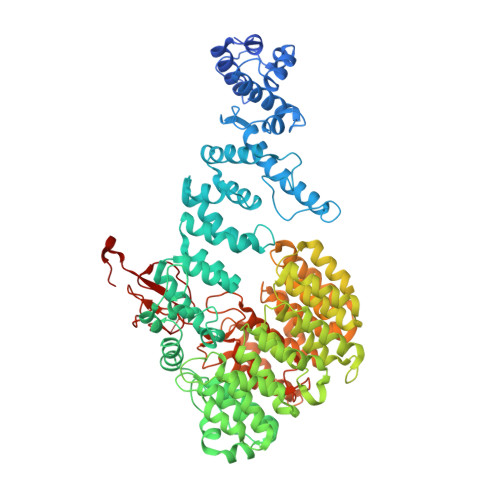
Structural Characterization of the Interaction of Ubp6 with the 26S Proteasome.
Aufderheide, A., Beck, F., Stengel, F., Hartwig, M., Schweitzer, A., Pfeifer, G., Goldberg, A.L., Sakata, E., Baumeister, W., Forster, F.(2015) Proc Natl Acad Sci U S A 112: 8626
- PubMed: 26130806 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1510449112
- Primary Citation Related Structures:
5A5B - PubMed Abstract:
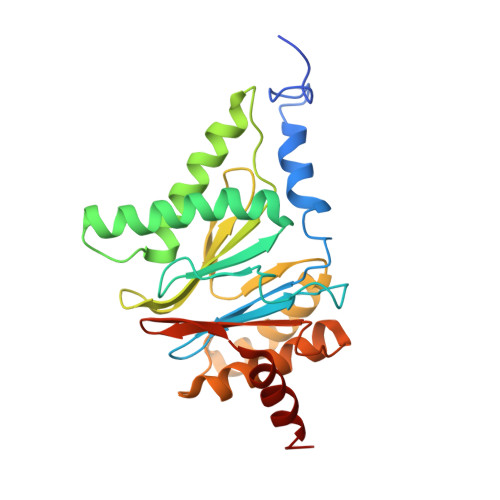
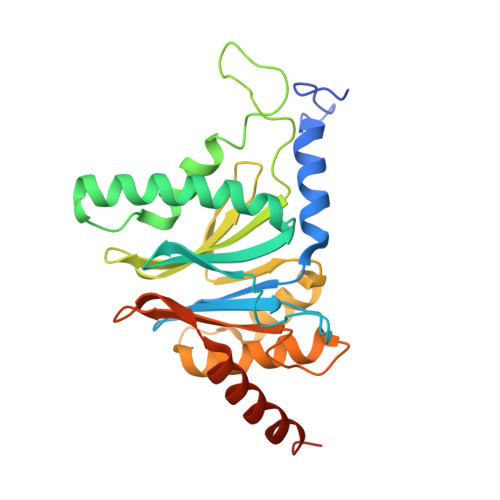
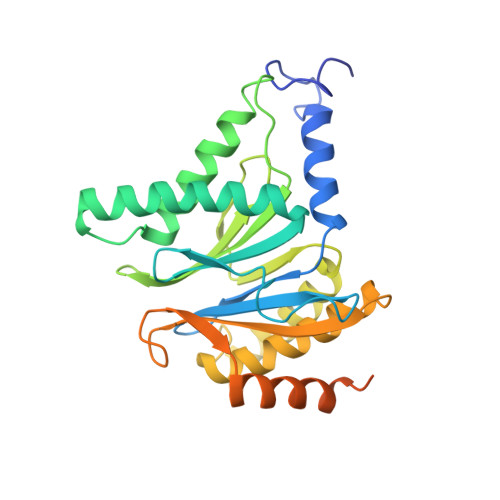
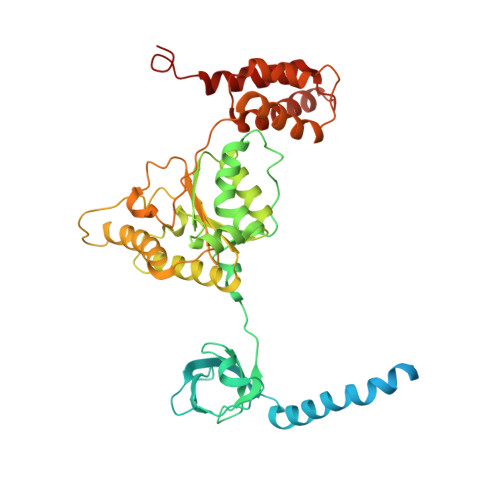
In eukaryotic cells, the 26S proteasome is responsible for the regulated degradation of intracellular proteins. Several cofactors interact transiently with this large macromolecular machine and modulate its function. The deubiquitylating enzyme ubiquitin C-terminal hydrolase 6 [Ubp6; ubiquitin-specific protease (USP) 14 in mammals] is the most abundant proteasome-interacting protein and has multiple roles in regulating proteasome function. Here, we investigate the structural basis of the interaction between Ubp6 and the 26S proteasome in the presence and absence of the inhibitor ubiquitin aldehyde. To this end we have used single-particle electron cryomicroscopy in combination with cross-linking and mass spectrometry. Ubp6 binds to the regulatory particle non-ATPase (Rpn) 1 via its N-terminal ubiquitin-like domain, whereas its catalytic USP domain is positioned variably. Addition of ubiquitin aldehyde stabilizes the binding of the USP domain in a position where it bridges the proteasome subunits Rpn1 and the regulatory particle triple-A ATPase (Rpt) 1. The USP domain binds to Rpt1 in the immediate vicinity of the Ubp6 active site, which may effect its activation. The catalytic triad is positioned in proximity to the mouth of the ATPase module and to the deubiquitylating enzyme Rpn11, strongly implying their functional linkage. On the proteasome side, binding of Ubp6 favors conformational switching of the 26S proteasome into an intermediate-energy conformational state, in particular upon the addition of ubiquitin aldehyde. This modulation of the conformational space of the 26S proteasome by Ubp6 explains the effects of Ubp6 on the kinetics of proteasomal degradation.
- Department of Molecular Structural Biology, Max Planck Institute of Biochemistry, 82152 Martinsried, Germany;
Organizational Affiliation: