Structural insights reveal the specific recognition of roX RNA by the dsRNA-binding domains of the RNA helicase MLE and its indispensable role in dosage compensation in Drosophila.
Lv, M., Yao, Y., Li, F., Xu, L., Yang, L., Gong, Q., Xu, Y.Z., Shi, Y., Fan, Y.J., Tang, Y.(2019) Nucleic Acids Res 47: 3142-3157
- PubMed: 30649456 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gky1308
- Primary Citation Related Structures:
5ZTM - PubMed Abstract:
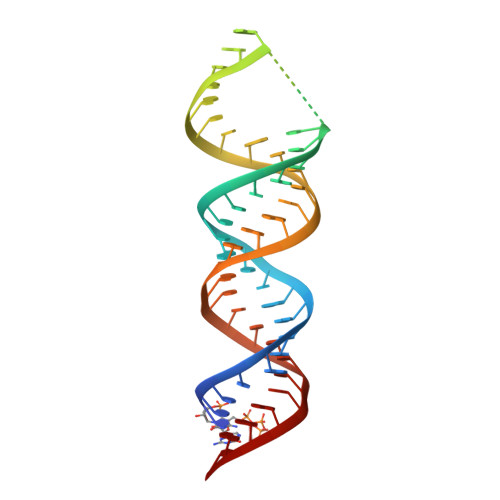
In Drosophila, dosage compensation globally upregulates the expression of genes located on male single X-chromosome. Maleless (MLE) helicase plays an essential role to incorporate the roX lncRNA into the dosage compensation complex (MSL-DCC), and such function is essentially dependent on its dsRNA-binding domains (dsRBDs). Here, we report a 2.90Å crystal structure of tandem dsRBDs of MLE in complex with a 55mer stem-loop of roX2 (R2H1). MLE dsRBDs bind to R2H1 cooperatively and interact with two successive minor grooves and a major groove of R2H1, respectively. The recognition of R2H1 by MLE dsRBDs involves both shape- and sequence-specificity. Moreover, dsRBD2 displays a stronger RNA affinity than dsRBD1, and mutations of key residues in either MLE dsRBD remarkably reduce their affinities for roX2 both in vitro and in vivo. In Drosophila, the structure-based mle mutations generated using the CRISPR/Cas9 system, are partially male-lethal and indicate the inter-regulation among the components of the MSL-DCC at multiple levels. Hence, our research provides structural insights into the interactions between MLE dsRBDs and R2H1 and facilitates a deeper understanding of the mechanism by which MLE tandem dsRBDs play an indispensable role in specific recognition of roX and the assembly of the MSL-DCC in Drosophila dosage compensation.
- Hefei National Laboratory for Physical Sciences at Microscale and School of Life Sciences, University of Science and Technology of China, Hefei, Anhui 230026, China.
Organizational Affiliation: