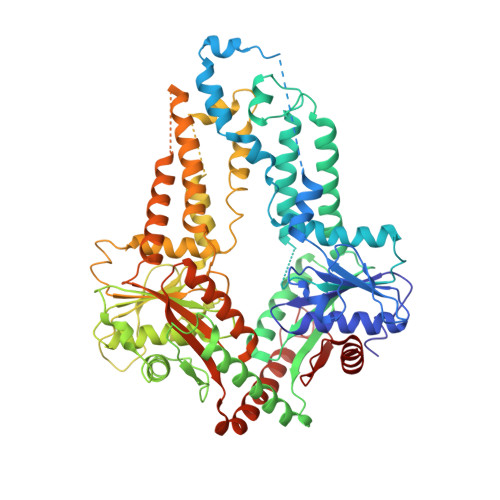
Structural analysis of chloroplast tail-anchored membrane protein recognition by ArsA1.
Lin, T.W., Chen, C.C., Wu, S.M., Chang, Y.C., Li, Y.C., Su, Y.W., Hsiao, C.D., Chang, H.Y.(2019) Plant J 99: 128-143
- PubMed: 30891827 Search on PubMed
- DOI: https://doi.org/10.1111/tpj.14316
- Primary Citation Related Structures:
5ZME, 5ZMF - PubMed Abstract:
In mammals and yeast, tail-anchored (TA) membrane proteins destined for the post-translational pathway are safely delivered to the endoplasmic reticulum (ER) membrane by a well-known targeting factor, TRC40/Get3. In contrast, the underlying mechanism for translocation of TA proteins in plants remains obscure. How this unique eukaryotic membrane-trafficking system correctly distinguishes different subsets of TA proteins destined for various organelles, including mitochondria, chloroplasts and the ER, is a key question of long standing. Here, we present crystal structures of algal ArsA1 (the Get3 homolog) in a distinct nucleotide-free open state and bound to adenylyl-imidodiphosphate. This approximately 80-kDa protein possesses a monomeric architecture, with two ATPase domains in a single polypeptide chain. It is capable of binding chloroplast (TOC34 and TOC159) and mitochondrial (TOM7) TA proteins based on features of its transmembrane domain as well as the regions immediately before and after the transmembrane domain. Several helices located above the TA-binding groove comprise the interlocking hook-like motif implicated by mutational analyses in TA substrate recognition. Our data provide insights into the molecular basis of the highly specific selectivity of interactions of algal ArsA1 with the correct sets of TA substrates before membrane targeting in plant cells.
- Molecular and Cell Biology, International Graduate Program, Academia Sinica and Graduate Institute of Life Science, National Defense Medical Center, Taipei, Taiwan.
Organizational Affiliation: