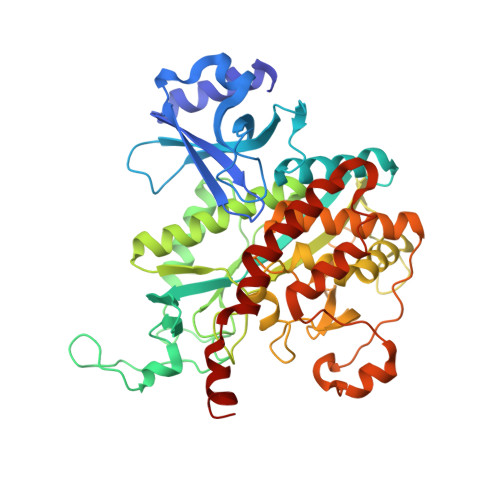
Structural Analysis of Glutamine Synthetase from Helicobacter pylori.
Joo, H.K., Park, Y.W., Jang, Y.Y., Lee, J.Y.(2018) Sci Rep 8: 11657-11657
- PubMed: 30076387 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-30191-5
- Primary Citation Related Structures:
5ZLI, 5ZLP - PubMed Abstract:
Glutamine synthetase (GS) is an enzyme that regulates nitrogen metabolism and synthesizes glutamine via glutamate, ATP, and ammonia. GS is a homo-oligomeric protein of eight, ten, or twelve subunits, and each subunit-subunit interface has its own active site. GS can be divided into GS I, GS II, and GS III. GS I and GS III form dodecamer in bacteria and archaea, whereas GS II form decamer in eukaryotes. GS I can be further subdivided into GS I-α and GS I-β according to its sequence and regulatory mechanism. GS is an essential protein for the survival of Helicobacter pylori which its infection could promote gastroduodenal diseases. Here, we determined the crystal structures of the GS from H. pylori (Hpy GS) in its apo- and substrate-bound forms at 2.8 Å and 2.9 Å resolution, respectively. Hpy GS formed a dodecamer composed of two hexameric rings stacked face-to-face. Hpy GS, which belongs to GS I, cannot be clearly classified as either GS I-α or GS I-β based on its sequence and regulatory mechanism. In this study, we propose that Hpy GS could be classified as a new GS-I subfamily and provide structural information on the apo- and substrate-bound forms of the protein.
- Department of Life Science, Dongguk University-Seoul, Ilsandong-gu, Goyang-si, Gyeonggi-do, 10326, Republic of Korea.
Organizational Affiliation: