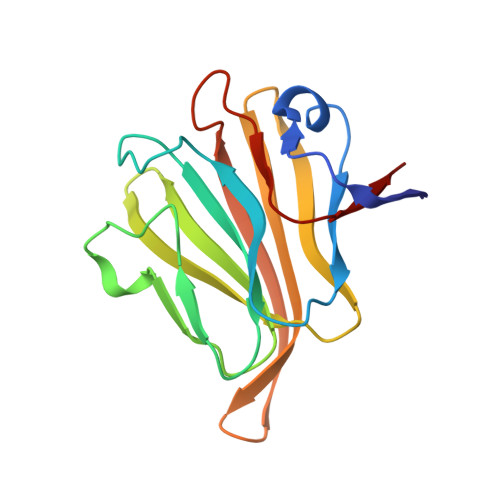
Human Group C Rotavirus VP8*s Recognize Type A Histo-Blood Group Antigens as Ligands.
Sun, X., Wang, L., Qi, J., Li, D., Wang, M., Cong, X., Peng, R., Chai, W., Zhang, Q., Wang, H., Wen, H., Gao, G.F., Tan, M., Duan, Z.(2018) J Virol 92
- PubMed: 29593033 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.00442-18
- Primary Citation Related Structures:
5ZHG, 5ZHO - PubMed Abstract:
Group/species C rotaviruses (RVCs) have been identified as important pathogens of acute gastroenteritis (AGE) in children, family-based outbreaks, as well as animal infections. However, little is known regarding their host-specific interaction, infection, and pathogenesis. In this study, we performed serial studies to characterize the function and structural features of a human G4P[2] RVC VP8* that is responsible for the host receptor interaction. Glycan microarrays demonstrated that the human RVC VP8* recognizes type A histo-blood group antigens (HBGAs), which was confirmed by synthetic glycan-/saliva-based binding assays and hemagglutination of red blood cells, establishing a paradigm of RVC VP8*-glycan interactions. Furthermore, the high-resolution crystal structure of the human RVC VP8* was solved, showing a typical galectin-like structure consisting of two β-sheets but with significant differences from cogent proteins of group A rotaviruses (RVAs). The VP8* in complex with a type A trisaccharide displays a novel ligand binding site that consists of a particular set of amino acid residues of the C-D, G-H, and K-L loops. RVC VP8* interacts with type A HBGAs through a unique mechanism compared with that used by RVAs. Our findings shed light on the host-virus interaction and the coevolution of RVCs and will facilitate the development of specific antivirals and vaccines. IMPORTANCE Group/species C rotaviruses (RVCs), members of Reoviridae family, infect both humans and animals, but our knowledge about the host factors that control host susceptibility and specificity is rudimentary. In this work, we characterized the glycan binding specificity and structural basis of a human RVC that recognizes type A HBGAs. We found that human RVC VP8*, the rotavirus host ligand binding domain that shares only ∼15% homology with the VP8* domains of RVAs, recognizes type A HBGA at an as-yet-unknown glycan binding site through a mechanism distinct from that used by RVAs. Our new advancements provide insights into RVC-cell attachment, the critical step of virus infection, which will in turn help the development of control and prevention strategies against RVs.
- Key Laboratory of Medical Virology and Viral Diseases, Ministry of Health of the People's Republic of China, Beijing, China.
Organizational Affiliation: