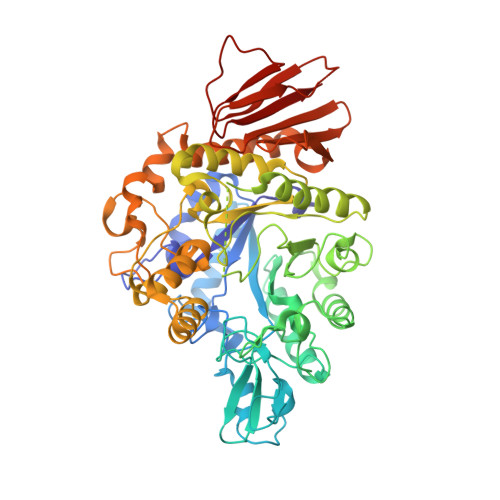
Function and structure of GH13_31 alpha-glucosidase with high alpha-(1→4)-glucosidic linkage specificity and transglucosylation activity.
Auiewiriyanukul, W., Saburi, W., Kato, K., Yao, M., Mori, H.(2018) FEBS Lett 592: 2268-2281
- PubMed: 29870070 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.13126
- Primary Citation Related Structures:
5ZCB, 5ZCC, 5ZCD, 5ZCE - PubMed Abstract:
α-Glucosidase hydrolyzes α-glucosides and transfers α-glucosyl residues to an acceptor through transglucosylation. In this study, GH13_31 α-glucosidase BspAG13_31A with high transglucosylation activity is reported in Bacillus sp. AHU2216 and biochemically and structurally characterized. This enzyme is specific to α-(1→4)-glucosidic linkage as substrates and transglucosylation products. Maltose is the most preferred substrate. Crystal structures of BspAG13_31A wild-type for the substrate-free form and inactive acid/base mutant E256Q in complexes with maltooligosaccharides were solved at 1.6-2.5 Å resolution. BspAG13_31A has a catalytic domain folded by an (β/α) 8 -barrel. In subsite +1, Ala200 and His203 on β→α loop 4 and Asn258 on β→α loop 5 are involved in the recognition of maltooligosaccharides. Structural basis for specificity of GH13_31 enzymes to α-(1→4)-glucosidic linkage is first described.
- Research Faculty of Agriculture, Hokkaido University, Sapporo, Japan.
Organizational Affiliation: