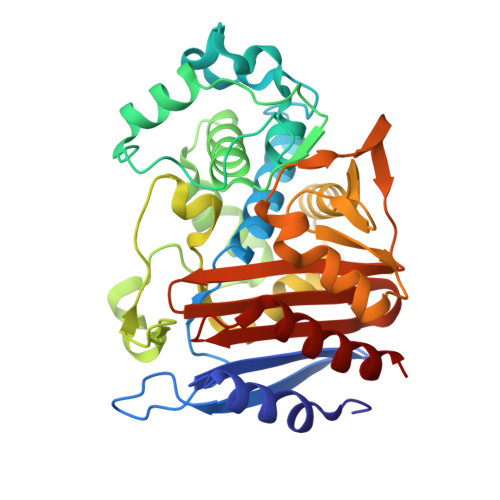
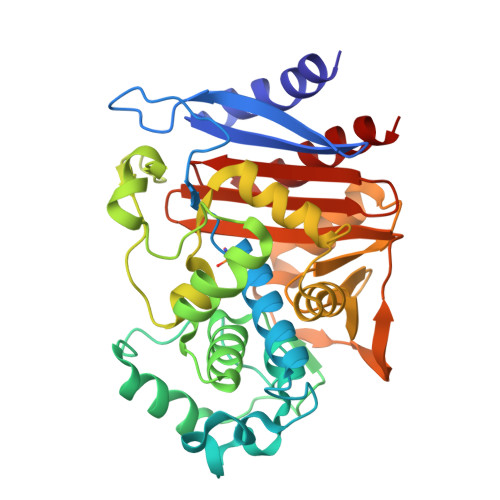
Probing the Mechanism of Inactivation of the FOX-4 Cephamycinase by Avibactam
Nukaga, M., Papp-Wallace, K.M., Hoshino, T., Lefurgy, S.T., Bethel, C.R., Barnes, M.D., Zeiser, E.T., Johnson, J.K., Bonomo, R.A.(2018) Antimicrob Agents Chemother 62
- PubMed: 29439972 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/AAC.02371-17
- Primary Citation Related Structures:
5ZA2 - PubMed Abstract:
Ceftazidime-avibactam is a "second-generation" β-lactam-β-lactamase inhibitor combination that is effective against Enterobacteriaceae expressing class A extended-spectrum β-lactamases, class A carbapenemases, and/or class C cephalosporinases. Knowledge of the interactions of avibactam, a diazabicyclooctane with different β-lactamases, is required to anticipate future resistance threats. FOX family β-lactamases possess unique hydrolytic properties with a broadened substrate profile to include cephamycins, partly as a result of an isoleucine at position 346, instead of the conserved asparagine found in most AmpCs. Interestingly, a single amino acid substitution at N346 in the Citrobacter AmpC is implicated in resistance to the aztreonam-avibactam combination. In order to understand how diverse active-site topologies affect avibactam inhibition, we tested a panel of clinical Enterobacteriaceae isolates producing bla FOX using ceftazidime-avibactam, determined the biochemical parameters for inhibition using the FOX-4 variant, and probed the atomic structure of avibactam with FOX-4. Avibactam restored susceptibility to ceftazidime for most isolates producing bla FOX ; two isolates, one expressing bla FOX-4 and the other producing bla FOX-5 , displayed an MIC of 16 μg/ml for the combination. FOX-4 possessed a k 2 / K value of 1,800 ± 100 M -1 · s -1 and an off rate ( k off ) of 0.0013 ± 0.0003 s -1 Mass spectrometry showed that the FOX-4-avibactam complex did not undergo chemical modification for 24 h. Analysis of the crystal structure of FOX-4 with avibactam at a 1.5-Å resolution revealed a unique characteristic of this AmpC β-lactamase. Unlike in the Pseudomonas -derived cephalosporinase 1 (PDC-1)-avibactam crystal structure, interactions (e.g., hydrogen bonding) between avibactam and position I346 in FOX-4 are not evident. Furthermore, another residue is not observed to be close enough to compensate for the loss of these critical hydrogen-bonding interactions. This observation supports findings from the inhibition analysis of FOX-4; FOX-4 possessed the highest K d (dissociation constant) value (1,600 nM) for avibactam compared to other AmpCs (7 to 660 nM). Medicinal chemists must consider the properties of extended-spectrum AmpCs, such as the FOX β-lactamases, for the design of future diazabicyclooctanes.
- Department of Pharmaceutical Sciences, Josai International University, Togane City, Chiba, Japan.
Organizational Affiliation: