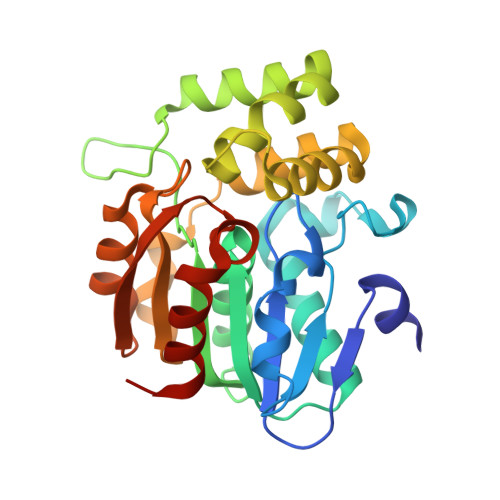
A missense allele of KARRIKIN-INSENSITIVE2 impairs ligand-binding and downstream signaling in Arabidopsis thaliana.
Lee, I., Kim, K., Lee, S., Lee, S., Hwang, E., Shin, K., Kim, D., Choi, J., Choi, H., Cha, J.S., Kim, H., Lee, R.A., Jeong, S., Kim, J., Kim, Y., Nam, H.G., Park, S.K., Cho, H.S., Soh, M.S.(2018) J Exp Bot 69: 3609-3623
- PubMed: 29722815 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/jxb/ery164
- Primary Citation Related Structures:
5Z9G, 5Z9H - PubMed Abstract:
A smoke-derived compound, karrikin (KAR), and an endogenous but as yet unidentified KARRIKIN INSENSITIVE2 (KAI2) ligand (KL) have been identified as chemical cues in higher plants that impact on multiple aspects of growth and development. Genetic screening of light-signaling mutants in Arabidopsis thaliana has identified a mutant designated as ply2 (pleiotropic long hypocotyl2) that has pleiotropic light-response defects. In this study, we used positional cloning to identify the molecular lesion of ply2 as a missense mutation of KAI2/HYPOSENSITIVE TO LIGHT, which causes a single amino acid substitution, Ala219Val. Physiological analysis and genetic epistasis analysis with the KL-signaling components MORE AXILLARY GROWTH2 (MAX2) and SUPPRESSOR OF MAX2 1 suggested that the pleiotropic phenotypes of the ply2 mutant can be ascribed to a defect in KL-signaling. Molecular and biochemical analyses revealed that the mutant KAI2ply2 protein is impaired in its ligand-binding activity. In support of this conclusion, X-ray crystallography studies suggested that the KAI2ply2 mutation not only results in a narrowed entrance gate for the ligand but also alters the structural flexibility of the helical lid domains. We discuss the structural implications of the Ala219 residue with regard to ligand-specific binding and signaling of KAI2, together with potential functions of KL-signaling in the context of the light-regulatory network in Arabidopsis thaliana.
- Division of Integrative Bioscience and Biotechnology, School of Life Science, Sejong University, Seoul, Republic of Korea.
Organizational Affiliation: