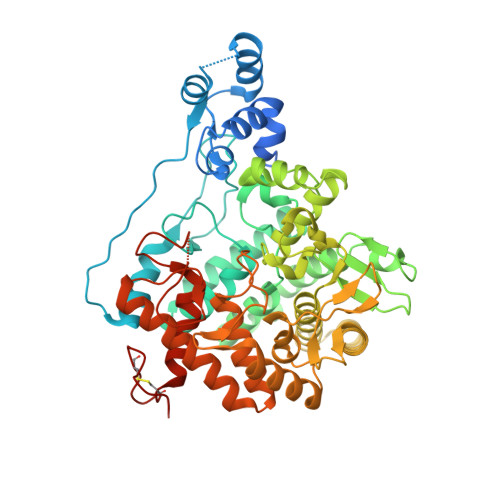
Mechanistic insights into enzymatic catalysis by trehalase from the insect gut endosymbiont Enterobacter cloacae.
Adhav, A., Harne, S., Bhide, A., Giri, A., Gayathri, P., Joshi, R.(2019) FEBS J 286: 1700-1716
- PubMed: 30657252 Search on PubMed
- DOI: https://doi.org/10.1111/febs.14760
- Primary Citation Related Structures:
5Z66, 5Z6H - PubMed Abstract:
Energy metabolism in the diamondback moth Plutella xylostella is facilitated by trehalase, an enzyme which assists in trehalose hydrolysis, from the predominant gut bacterium Enterobacter cloacae. We report the biochemical and structural characterization of recombinant trehalase from E. cloacae (Px_EclTre). Px_EclTre showed K M of 1.47 (±0.05) mm, k cat of 6254.72 min -1 and V max 0.2 (±0.002) mm·min -1 at 55 °C and acidic pH. Crystal structures of Px_EclTre were determined in the ligand-free form and bound to the inhibitor Validoxylamine A. The crystal structure of the ligand-free form, unavailable until now for any other bacterial trehalases, enabled us to delineate the conformational changes accompanying ligand binding in trehalases. Multiple salt bridges were identified that potentially facilitated closure of a hood over the substrate-binding site. A cluster of five tryptophans lined the -1 substrate-binding subsite, interacted with crucial active site residues and contributed to both trehalase activity and stability. The importance of these residues in enzyme activity was further validated by mutagenesis studies. Many of these identified residues form part of signature motifs and other conserved sequences in trehalases. The structure analysis thus led to the assignment of the functional role to these conserved residues. This information can be further explored for the design of effective inhibitors against trehalases.
- Institute of Bioinformatics and Biotechnology, Savitribai Phule Pune University, India.
Organizational Affiliation: