Structural insights into the interaction of helicase and primase inMycobacterium tuberculosis.
Sharma, D.P., Vijayan, R., Rehman, S.A.A., Gourinath, S.(2018) Biochem J 475: 3493-3509
- PubMed: 30315069 Search on PubMed
- DOI: https://doi.org/10.1042/BCJ20180673
- Primary Citation Related Structures:
5Z51 - PubMed Abstract:
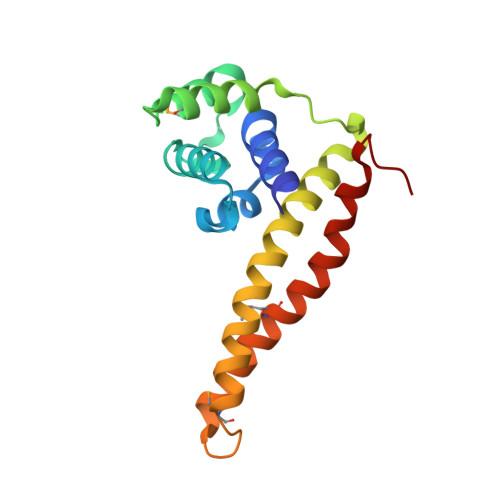
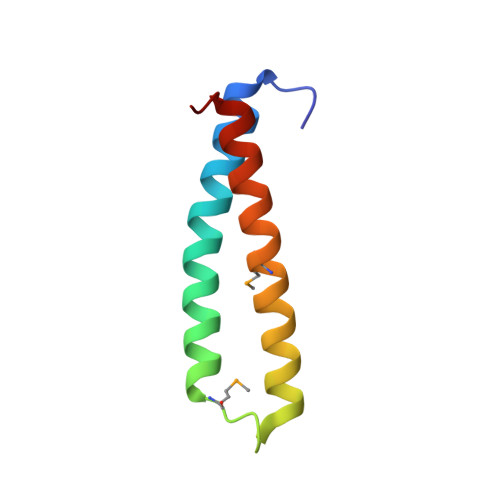
The helicase-primase interaction is an essential event in DNA replication and is mediated by the highly variable C-terminal domain of primase (DnaG) and N-terminal domain of helicase (DnaB). To understand the functional conservation despite the low sequence homology of the DnaB-binding domains of DnaGs of eubacteria, we determined the crystal structure of the helicase-binding domain of DnaG from Mycobacterium tuberculosis ( Mt DnaG-CTD) and did so to a resolution of 1.58 Å. We observed the overall structure of Mt DnaG-CTD to consist of two subdomains, the N-terminal globular region (GR) and the C-terminal helical hairpin region (HHR), connected by a small loop. Despite differences in some of its helices, the globular region was found to have broadly similar arrangements across the species, whereas the helical hairpins showed different orientations. To gain insights into the crucial helicase-primase interaction in M. tuberculosis , a complex was modeled using the Mt DnaG-CTD and Mt DnaB-NTD crystal structures. Two nonconserved hydrophobic residues (Ile605 and Phe615) of Mt DnaG were identified as potential key residues interacting with Mt DnaB. Biosensor-binding studies showed a significant decrease in the binding affinity of Mt DnaB-NTD with the Ile605Ala mutant of Mt DnaG-CTD compared with native Mt DnaG-CTD. The loop, connecting the two helices of the HHR, was concluded to be largely responsible for the stability of the DnaB-DnaG complex. Also, Mt DnaB-NTD showed micromolar affinity with DnaG-CTDs from Escherichia coli and Helicobacter pylori and unstable binding with DnaG-CTD from Vibrio cholerae The interacting domains of both DnaG and DnaB demonstrate the species-specific evolution of the replication initiation system.
- School of Life Sciences, Jawaharlal Nehru University, Delhi 110067, India.
Organizational Affiliation: