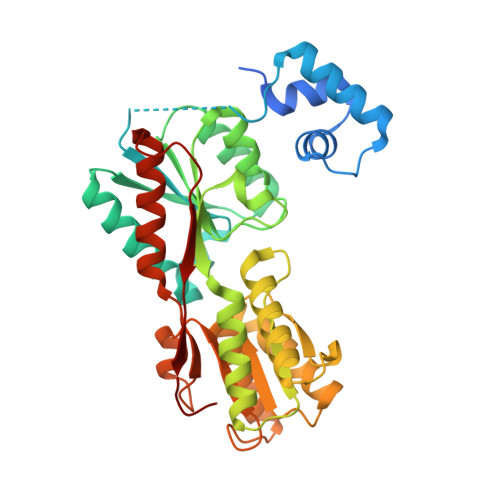
Structural and functional analyses of the cellulase transcription regulator CelR
Fu, Y., Yeom, S.J., Kwon, K.K., Hwang, J., Kim, H., Woo, E.J., Lee, D.H., Lee, S.G.(2018) FEBS Lett 592: 2776-2785
- PubMed: 30062758 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.13206
- Primary Citation Related Structures:
5YSZ - PubMed Abstract:
CelR is a transcriptional regulator that controls the expression of cellulases catalyzing cellulose hydrolysis. However, the structural mechanism of its regulation has remained unclear. Here, we report the first structure of CelR, in this case with cellobiose bound. CelR consists of a DNA-binding domain (DBD) and a regulatory domain (RD), and homodimerizes with each monomer bound to cellobiose. A hinge region (HR) in CelR connects the DBD with the RD, and Leu59 in the HR acts as a 'leucine lever' that transduces a transcriptional activation signal. Furthermore, an α4 helix mediates the ligand-binding signal for transcriptional activation. Tyr84 and Gln301 can potentially alter the ligand specificity of CelR. This study provides a pivotal step toward understanding transcription of the cellulases.
- Synthetic Biology and Bioengineering Research Center, Korea Research Institute of Bioscience and Biotechnology (KRIBB), Daejeon, Korea.
Organizational Affiliation: