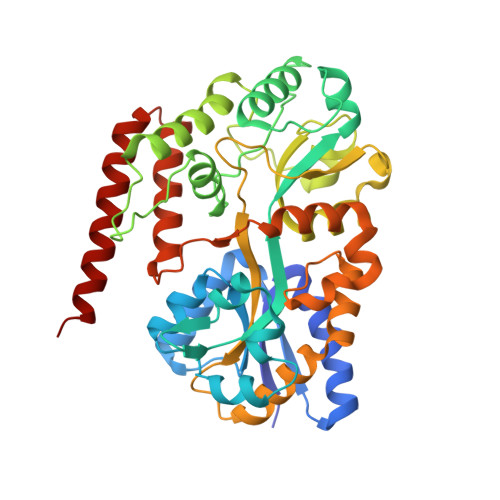
Structural and thermodynamic insights into beta-1,2-glucooligosaccharide capture by a solute-binding protein inListeria innocua.
Abe, K., Sunagawa, N., Terada, T., Takahashi, Y., Arakawa, T., Igarashi, K., Samejima, M., Nakai, H., Taguchi, H., Nakajima, M., Fushinobu, S.(2018) J Biological Chem 293: 8812-8828
- PubMed: 29678880 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA117.001536
- Primary Citation Related Structures:
5YSB, 5YSD, 5YSE, 5YSF - PubMed Abstract:
β-1,2-Glucans are bacterial carbohydrates that exist in cyclic or linear forms and play an important role in infections and symbioses involving Gram-negative bacteria. Although several β-1,2-glucan-associated enzymes have been characterized, little is known about how β-1,2-glucan and its shorter oligosaccharides (Sop n s) are captured and imported into the bacterial cell. Here, we report the biochemical and structural characteristics of the Sop n -binding protein (SO-BP, Lin1841) associated with the ATP-binding cassette (ABC) transporter from the Gram-positive bacterium Listeria innocua Calorimetric analysis revealed that SO-BP specifically binds to Sop n s with a degree of polymerization of 3 or more, with K d values in the micromolar range. The crystal structures of SO-BP in an unliganded open form and in closed complexes with tri-, tetra-, and pentaoligosaccharides (Sop 3-5 ) were determined to a maximum resolution of 1.6 Å. The binding site displayed shape complementarity to Sop n , which adopted a zigzag conformation. We noted that water-mediated hydrogen bonds and stacking interactions play a pivotal role in the recognition of Sop 3-5 by SO-BP, consistent with its binding thermodynamics. Computational free-energy calculations and a mutational analysis confirmed that interactions with the third glucose moiety of Sop n s are significantly responsible for ligand binding. A reduction in unfavorable changes in binding entropy that were in proportion to the lengths of the Sop n s was explained by conformational entropy changes. Phylogenetic and sequence analyses indicated that SO-BP ABC transporter homologs, glycoside hydrolases, and other related proteins are co-localized in the genomes of several bacteria. This study may improve our understanding of bacterial β-1,2-glucan metabolism and promote the discovery of unidentified β-1,2-glucan-associated proteins.
- From the Department of Biotechnology.
Organizational Affiliation: