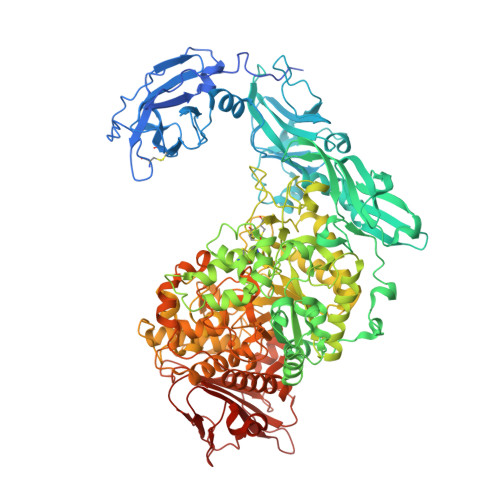
Elucidation of the mechanism of interaction between Klebsiella pneumoniae pullulanase and cyclodextrin
Saka, N., Iwamoto, H., Malle, D., Takahashi, N., Mizutani, K., Mikami, B.(2018) Acta Crystallogr D Struct Biol 74: 1115-1123
- PubMed: 30387770 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798318014523
- Primary Citation Related Structures:
5YN2, 5YN7, 5YNA, 5YNC, 5YND, 5YNE, 5YNH - PubMed Abstract:
Crystal structures of Klebsiella pneumoniae pullulanase (KPP) in complex with α-cyclodextrin (α-CD), β-cyclodextrin (β-CD) and γ-cyclodextrin (γ-CD) were refined at around 1.98-2.59 Å resolution from data collected at SPring-8. In the structures of the complexes obtained with 1 mM α-CD or γ-CD, one molecule of CD was found at carbohydrate-binding module 41 only (CBM41). In the structures of the complexes obtained with 1 mM β-CD or with 10 mM α-CD or γ-CD, two molecules of CD were found at CBM41 and in the active-site cleft, where the hydrophobic residue of Phe746 occupies the inside cavity of the CD rings. In contrast to α-CD and γ-CD, one β-CD molecule was found at the active site only in the presence of 0.1 mM β-CD. These results were coincident with the solution experiments, which showed that β-CD inhibits this enzyme more than a thousand times more potently than α-CD and γ-CD. The strong inhibition of β-CD is caused by the optimized interaction between β-CD and the side chain of Phe746. The increased K i values of the F746A mutant for β-CD supported the importance of Phe746 in the strong interaction of pullulanase with β-CD.
- Laboratory of Applied Structural Biology, Division of Applied Life Sciences, Graduate School of Agriculture, Kyoto University, Gokasyo, Uji, Kyoto 611-0011, Japan.
Organizational Affiliation: