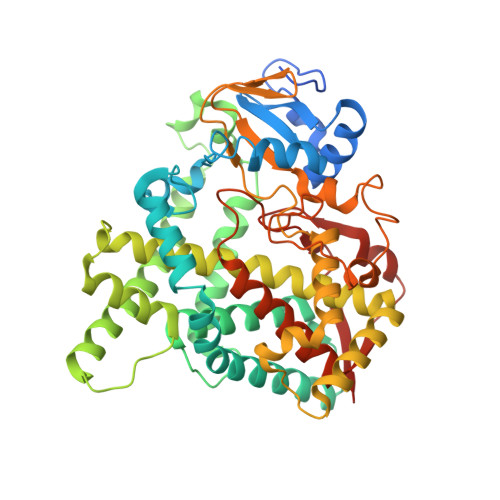
Crystal structure of CYP76AH1 in 4-PI-bound state from Salvia miltiorrhiza.
Gu, M., Wang, M., Guo, J., Shi, C., Deng, J., Huang, L., Huang, L., Chang, Z.(2019) Biochem Biophys Res Commun 511: 813-819
- PubMed: 30837155 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2019.02.103
- Primary Citation Related Structures:
5YM3 - PubMed Abstract:
Tanshinones are important diterpenoid secondary metabolites from Salvia miltiorrhiza, widely used as cardiovascular and cerebrovascular medicines. CYP76AH1 is a membrane-associated cytochrome P450 enzyme and plays a critical role in the biosynthetic pathway of tanshinones. To clarify the relationship between structure and function of CYP76AH1, we recently constructed the expression vector of CYP76AH1 and purified the enzyme. The engineered CYP76AH1 was expressed in E. coli Trans-blue cells and exhibited enhanced expression and solubility. The proper folding of the engineered CYP76AH1 was assessed by CO difference spectrum assay. Functional identification of the recombinant enzyme was performed by conducting enzymatic reaction with the purified CYP76AH1 in presence of substrate, the co-factor NADPH and the purified SmCPR1 (cytochrome P450 reductase from Salvia miltiorrhiza), and by subsequently analyzing the reaction extract through GC-MS. X-ray crystal complex structure of CYP76AH1 with inhibitor 4-phenylimmidazole (4-PI) was determined at the resolution of 2.6 Å. In the ligand-binding cavity of 4-PI bound CYP76AH1, the inhibitor 4-PI forms a hydrogen bound with a water molecule which coordinates with heme at the sixth coordination position. There are two open channels which substrate and product site may access and leave the active site. In the CYP76AH1/4-PI complex structure, the imidazole ring of 4-PI is parallel to helix I instead of perpendicular to helies I in most P450s bound imidazole. 4-PI may be work in the stability of CYP76AH1 crystal structure. These studies provide information on functional expression and purification of CYP76AH1, and overall structure of CYP76AH1 complexed with 4-PI.
- Department of Biophysics, School of Basic Medical Sciences, Peking University, Beijing, 100191, China.
Organizational Affiliation: