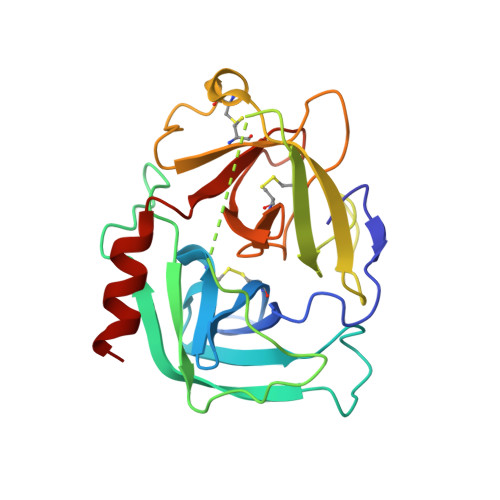
Structure-based design, synthesis, and binding mode analysis of novel and potent chymase inhibitors
Futamura-Takahashi, J., Tanaka, T., Sugawara, H., Iwashita, S., Imajo, S., Oyama, Y., Muto, T.(2018) Bioorg Med Chem Lett 28: 188-192
- PubMed: 29191554 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2017.11.031
- Primary Citation Related Structures:
5YJM, 5YJP - PubMed Abstract:
Based on insight from the X-ray crystal structure of human chymase in complex with compound 1, a lactam carbonyl of the diazepane core was exchanged with O-substituted oxyimino group, leading to amidoxime derivatives. This modification resulted in highly potent chymase inhibitors, such as O-phenylamidoxime 5f. X-ray crystal structure analysis indicated that compound 5f induced movement of the Leu99 and Tyr94 side chains at the S2 site, and the increase in inhibitory activity of O-phenyl amidoxime derivatives suggested that the O-phenyl moiety interacted with the Tyr94 residue. Surface plasmon resonance experiments showed that compound 5f had slower association and dissociation kinetics and the calculated residence time of compound 5f to human chymase was extended compared to that of amide compound 1.
- Asubio Pharma Co., Ltd, 6-4-3 Minatojima-Minamimachi, Chuo-ku, Kobe 650-0047, Japan. Electronic address: futamura.junko.b6@asubio.co.jp.
Organizational Affiliation: