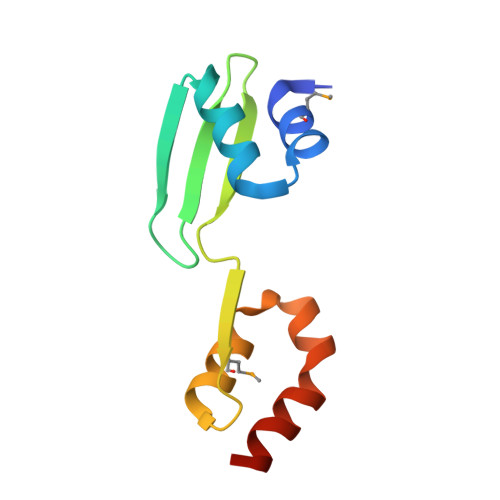
CRISPR RNA and anti-CRISPR protein binding to theXanthomonas albilineansCsy1-Csy2 heterodimer in the type I-F CRISPR-Cas system
Hong, S., Ka, D., Yoon, S.J., Suh, N., Jeong, M., Suh, J.Y., Bae, E.(2018) J Biological Chem 293: 2744-2754
- PubMed: 29348170 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA117.001611
- Primary Citation Related Structures:
5YHR - PubMed Abstract:
Clustered regularly interspaced short palindromic repeats (CRISPRs) and CRISPR-associated (Cas) proteins provide microbial adaptive immunity against bacteriophages. In type I-F CRISPR-Cas systems, multiple Cas proteins (Csy1-4) compose a surveillance complex (Csy complex) with CRISPR RNA (crRNA) for target recognition. Here, we report the biochemical characterization of the Csy1-Csy2 subcomplex from Xanthomonas albilineans , including the analysis of its interaction with crRNA and AcrF2, an anti-CRISPR (Acr) protein from a phage that infects Pseudomonas aeruginosa The X. albilineans Csy1 and Csy2 proteins (XaCsy1 and XaCsy2, respectively) formed a stable heterodimeric complex that specifically bound the 8-nucleotide (nt) 5'-handle of the crRNA. In contrast, the XaCsy1-XaCsy2 heterodimer exhibited reduced affinity for the 28-nt X. albilineans CRISPR repeat RNA containing the 5'-handle sequence. Chromatographic and calorimetric analyses revealed tight binding between the Acr protein from the P. aeruginosa phage and the heterodimeric subunit of the X. albilineans Csy complex, suggesting that AcrF2 recognizes conserved features of Csy1-Csy2 heterodimers. We found that neither XaCsy1 nor XaCsy2 alone forms a stable complex with AcrF2 and the 5'-handle RNA, indicating that XaCsy1-XaCsy2 heterodimerization is required for binding them. We also solved the crystal structure of AcrF2 to a resolution of 1.34 Å, enabling a more detailed structural analysis of the residues involved in the interactions with the Csy1-Csy2 heterodimer. Our results provide information about the order of events during the formation of the multisubunit crRNA-guided surveillance complex and suggest that the Acr protein inactivating type I-F CRISPR-Cas systems has broad specificity.
- Department of Agricultural Biotechnology, Seoul National University, Seoul 08826, Korea.
Organizational Affiliation: