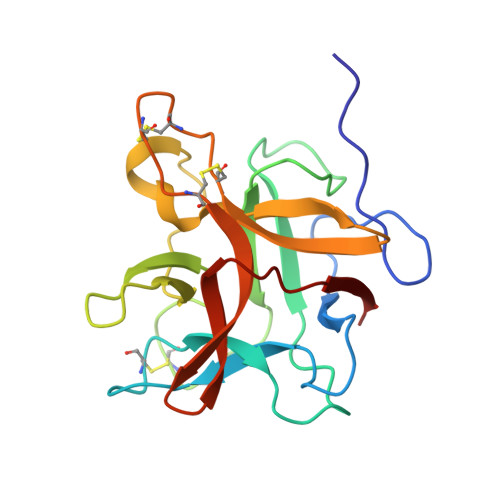
Structural and functional analysis of miraculin-like protein from Vitis vinifera.
Ohkura, S.I., Hori, M., Saitoh, K., Okuzawa, T., Okamoto, I., Furukawa, N., Shimizu-Ibuka, A.(2018) Biochim Biophys Acta Proteins Proteom 1866: 1125-1130
- PubMed: 30282610 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbapap.2018.08.009
- Primary Citation Related Structures:
5YH4 - PubMed Abstract:
The so-called miraculin-like proteins (MLPs) are homologous to miraculin, a homodimeric protein with taste-modifying activity that converts sourness into sweetness. The identity between MLPs and miraculin generally ranges from 30% to 55%, and both MLPs and miraculin are categorized into the Kunitz-type soybean trypsin inhibitor (STI) family. MLP from grape (Vitis vinifera; vvMLP) exhibits significant homology to miraculin (61% identity), suggesting that vvMLP possesses miraculin-like properties. The results of size-exclusion chromatography and sensory analysis illustrated that vvMLP exists as a monomer in solution with no detectable taste-modifying activity. Crystal structure determination revealed that vvMLP exists as a β-trefoil fold, similarly as other MLPs and Kunitz-type protein inhibitors. The conformation of the loops, including the so-called reactive loop in the STI family, was substantially different between vvMLP and STI. Recombinant vvMLP had inhibitory activity against trypsin (K i = 13.7 μM), indicating that the protein can act as a moderate trypsin inhibitor.
- Faculty of Applied Life Sciences, Niigata University of Pharmacy and Applied Life Sciences, 265-1 Higashijima, Akiha-ku, Niigata 956-8603, Japan.
Organizational Affiliation: