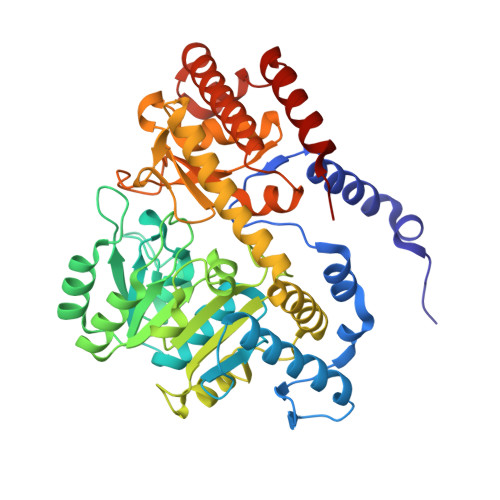
Potent Inhibitors of Plasmodial Serine Hydroxymethyltransferase (SHMT) Featuring a Spirocyclic Scaffold
Schwertz, G., Witschel, M.C., Rottmann, M., Leartsakulpanich, U., Chitnumsub, P., Jaruwat, A., Amornwatcharapong, W., Ittarat, W., Schafer, A., Aponte, R.A., Trapp, N., Chaiyen, P., Diederich, F.(2018) ChemMedChem 13: 931-943
- PubMed: 29655285 Search on PubMed
- DOI: https://doi.org/10.1002/cmdc.201800053
- Primary Citation Related Structures:
5YFZ, 5YG2, 5YG3, 5YG4 - PubMed Abstract:
With the discovery that serine hydroxymethyltransferase (SHMT) is a druggable target for antimalarials, the aim of this study was to design novel inhibitors of this key enzyme in the folate biosynthesis cycle. Herein, 19 novel spirocyclic ligands based on either 2-indolinone or dihydroindene scaffolds and featuring a pyrazolopyran core are reported. Strong target affinities for Plasmodium falciparum (Pf) SHMT (14-76 nm) and cellular potencies in the low nanomolar range (165-334 nm) were measured together with interesting selectivity against human cytosolic SHMT1 (hSHMT1). Four co-crystal structures with Plasmodium vivax (Pv) SHMT solved at 2.2-2.4 Å resolution revealed the key role of the vinylogous cyanamide for anchoring ligands within the active site. The spirocyclic motif in the molecules enforces the pyrazolopyran core to adopt a substantially more curved conformation than that of previous non-spirocyclic analogues. Finally, solvation of the spirocyclic lactam ring of the receptor-bound ligands is discussed.
- Laboratorium für Organische Chemie, ETH Zürich, Vladimir-Prelog-Weg 3, 8093, Zürich, Switzerland.
Organizational Affiliation: