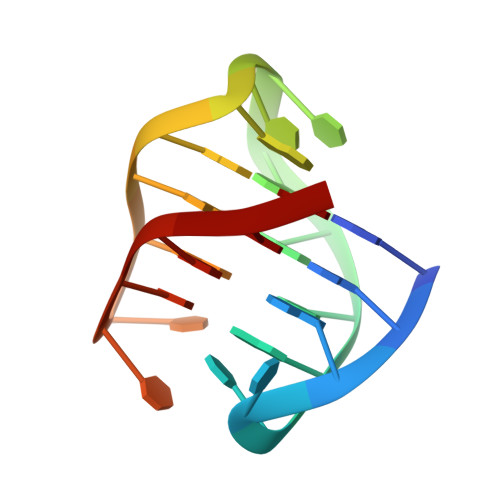
A chair-type G-quadruplex structure formed by a human telomeric variant DNA in K+solution.
Liu, C., Zhou, B., Geng, Y., Yan Tam, D., Feng, R., Miao, H., Xu, N., Shi, X., You, Y., Hong, Y., Tang, B.Z., Kwan Lo, P., Kuryavyi, V., Zhu, G.(2019) Chem Sci 10: 218-226
- PubMed: 30713633 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/c8sc03813a
- Primary Citation Related Structures:
5YEY - PubMed Abstract:
Guanine tracts of human telomeric DNA sequences are known to fold into eight different four-stranded structures that vary by the conformation of guanine nucleotides arranged in the stack of G-tetrads in their core and by different kinds and orders of connecting loops, called G-quadruplexes. Here, we present a novel G-quadruplex structure formed in K + solution by a human telomeric variant d[(GGGTTA)2GGGTTTGGG], htel21 T 18 . This variant DNA is located in the subtelomeric regions of human chromosomes 8, 11, 17, and 19 as well as in the DNase hypersensitive region and in the subcentromeric region of chromosome 5. Interestingly, single A18T substitution that makes htel21 T 18 different from the human telomeric sequence results in the formation of a three-layer chair-type G-quadruplex, a fold previously unknown among human telomeric repeats, with two loops interacting through the reverse Watson-Crick A6·T18 base pair. The loops are edgewise; glycosidic conformation of guanines is syn · anti · syn · anti around each tetrad, and each strand of the core has two antiparallel adjacent strands. Our results expand the repertoire of known G-quadruplex folding topologies and may provide a potential target for structure-based anticancer drug design.
- Division of Life Science , The Hong Kong University of Science and Technology , Clear Water Bay , Kowloon , Hong Kong SAR , China . Email: gzhu@ust.hk.
Organizational Affiliation: