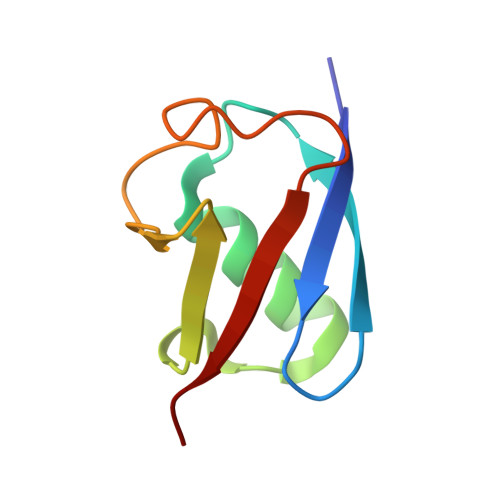
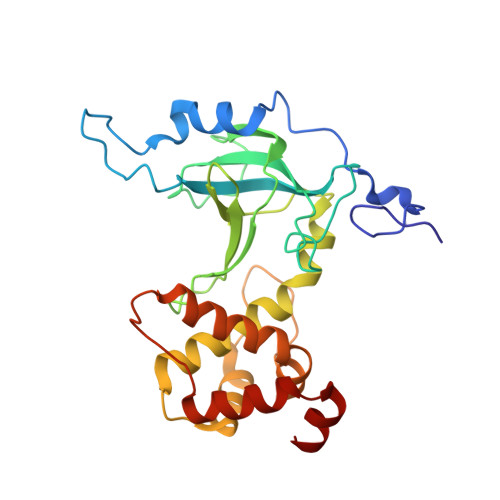
Structural and mechanistic insights into UHRF1-mediated DNMT1 activation in the maintenance DNA methylation.
Li, T., Wang, L., Du, Y., Xie, S., Yang, X., Lian, F., Zhou, Z., Qian, C.(2018) Nucleic Acids Res 46: 3218-3231
- PubMed: 29471350 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gky104
- Primary Citation Related Structures:
5YDR - PubMed Abstract:
UHRF1 plays multiple roles in regulating DNMT1-mediated DNA methylation maintenance during DNA replication. The UHRF1 C-terminal RING finger functions as an ubiquitin E3 ligase to establish histone H3 ubiquitination at Lys18 and/or Lys23, which is subsequently recognized by DNMT1 to promote its localization onto replication foci. Here, we present the crystal structure of DNMT1 RFTS domain in complex with ubiquitin and highlight a unique ubiquitin binding mode for the RFTS domain. We provide evidence that UHRF1 N-terminal ubiquitin-like domain (UBL) also binds directly to DNMT1. Despite sharing a high degree of structural similarity, UHRF1 UBL and ubiquitin bind to DNMT1 in a very distinct fashion and exert different impacts on DNMT1 enzymatic activity. We further show that the UHRF1 UBL-mediated interaction between UHRF1 and DNMT1, and the binding of DNMT1 to ubiquitinated histone H3 that is catalyzed by UHRF1 RING domain are critical for the proper subnuclear localization of DNMT1 and maintenance of DNA methylation. Collectively, our study adds another layer of complexity to the regulatory mechanism of DNMT1 activation by UHRF1 and supports that individual domains of UHRF1 participate and act in concert to maintain DNA methylation patterns.
- School of Biomedical Sciences, The University of Hong Kong, PokFuLam, Hong Kong.
Organizational Affiliation: