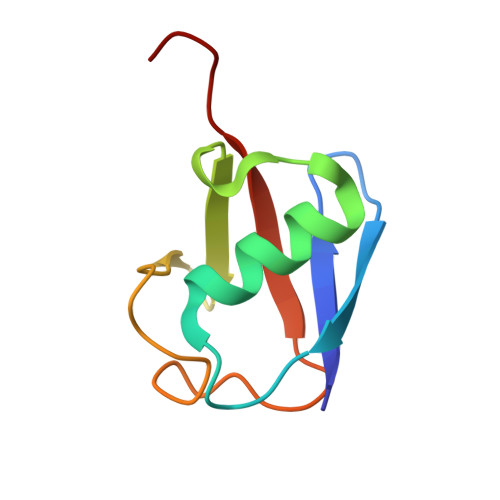
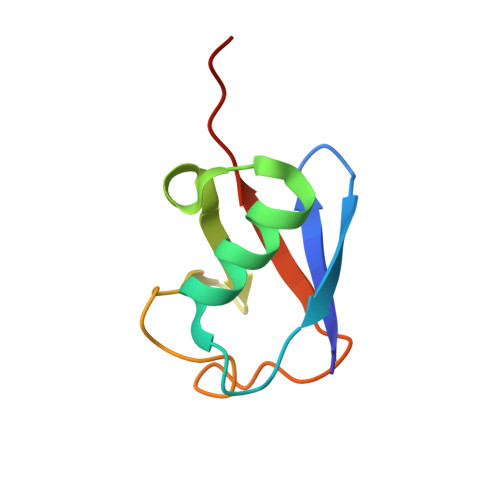
Structural insights into two distinct binding modules for Lys63-linked polyubiquitin chains in RNF168
Takahashi, T.S., Hirade, Y., Toma, A., Sato, Y., Yamagata, A., Goto-Ito, S., Tomita, A., Nakada, S., Fukai, S.(2018) Nat Commun 9: 170-170
- PubMed: 29330428 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-02345-y
- Primary Citation Related Structures:
5XIS, 5XIT, 5XIU, 5YDK - PubMed Abstract:
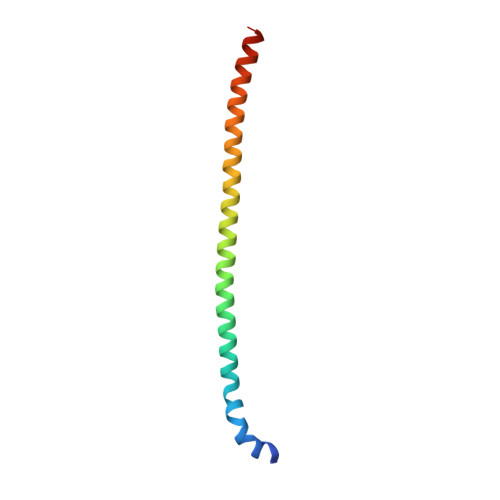
The E3 ubiquitin (Ub) ligase RNF168 plays a critical role in the initiation of the DNA damage response to double-strand breaks (DSBs). The recruitment of RNF168 by ubiquitylated targets involves two distinct regions, Ub-dependent DSB recruitment module (UDM) 1 and UDM2. Here we report the crystal structures of the complex between UDM1 and Lys63-linked diUb (K63-Ub 2 ) and that between the C-terminally truncated UDM2 (UDM2ΔC) and K63-Ub 2 . In both structures, UDM1 and UDM2ΔC fold as a single α-helix. Their simultaneous bindings to the distal and proximal Ub moieties provide specificity for Lys63-linked Ub chains. Structural and biochemical analyses of UDM1 elucidate an Ub-binding mechanism between UDM1 and polyubiquitylated targets. Mutations of Ub-interacting residues in UDM2 prevent the accumulation of RNF168 to DSB sites in U2OS cells, whereas those in UDM1 have little effect, suggesting that the interaction of UDM2 with ubiquitylated and polyubiquitylated targets mainly contributes to the RNF168 recruitment.
- Institute of Molecular and Cellular Biosciences, The University of Tokyo, Tokyo, 113-0032, Japan.
Organizational Affiliation: