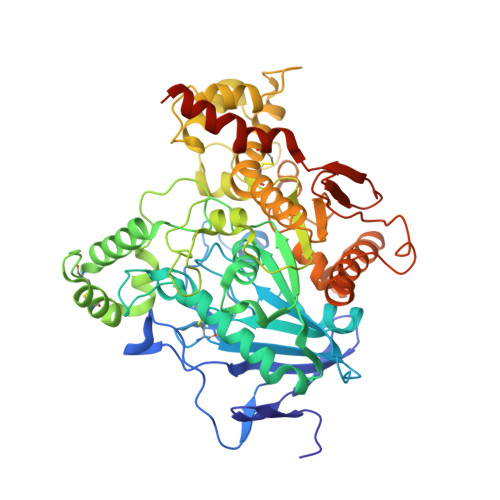
Crystal structures of acetylcholinesterase of the malaria vector Anopheles gambiae reveal a polymerization interface, ligand binding residues and post translational modifications
Han, Q., Guan, H., Ding, H., Liao, C., Robinson, H., Li, J.To be published.