Assessing the Heterogeneity of the Fc-Glycan of a Therapeutic Antibody Using an engineered Fc gamma Receptor IIIa-Immobilized Column.
Kiyoshi, M., Caaveiro, J.M.M., Tada, M., Tamura, H., Tanaka, T., Terao, Y., Morante, K., Harazono, A., Hashii, N., Shibata, H., Kuroda, D., Nagatoishi, S., Oe, S., Ide, T., Tsumoto, K., Ishii-Watabe, A.(2018) Sci Rep 8: 3955-3955
- PubMed: 29500371 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-22199-8
- Primary Citation Related Structures:
5YC5 - PubMed Abstract:
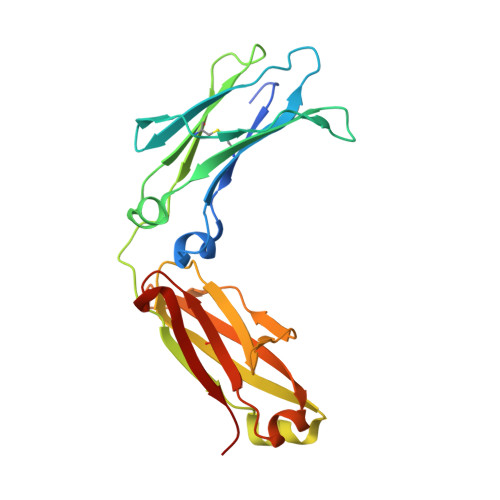
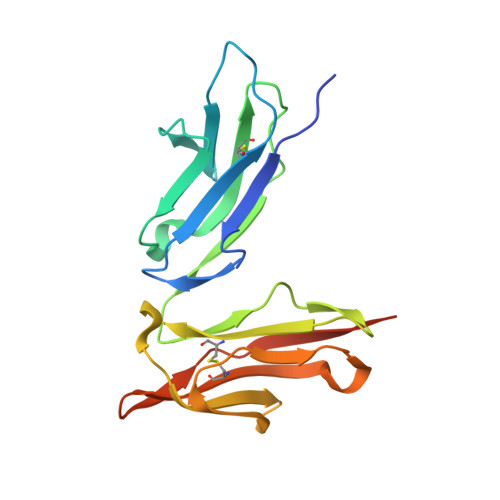
The N-glycan moiety of IgG-Fc has a significant impact on multifaceted properties of antibodies such as in their effector function, structure, and stability. Numerous studies have been devoted to understanding its biological effect since the exact composition of the Fc N-glycan modulates the magnitude of effector functions such as the antibody-dependent cell mediated cytotoxicity (ADCC), and the complement-dependent cytotoxicity (CDC). To date, systematic analyses of the properties and influence of glycan variants have been of great interest. Understanding the principles on how N-glycosylation modulates those properties is important for the molecular design, manufacturing, process optimization, and quality control of therapeutic antibodies. In this study, we have separated a model therapeutic antibody into three fractions according to the composition of the N-glycan by using a novel FcγRIIIa chromatography column. Notably, Fc galactosylation was a major factor influencing the affinity of IgG-Fc to the FcγRIIIa immobilized on the column. Each antibody fraction was employed for structural, biological, and physicochemical analysis, illustrating the mechanism by which galactose modulates the affinity to FcγRIIIa. In addition, we discuss the benefits of the FcγRIIIa chromatography column to assess the heterogeneity of the N-glycan.
- Division of Biological Chemistry and Biologicals, National Institute of Health Sciences, Tokyo, 158-8501, Japan. m.kiyoshi@nihs.go.jp.
Organizational Affiliation: