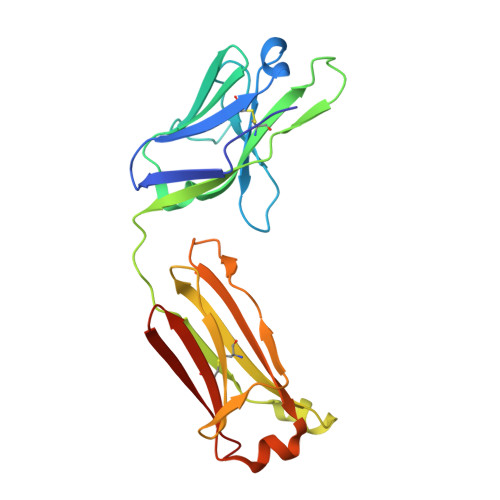
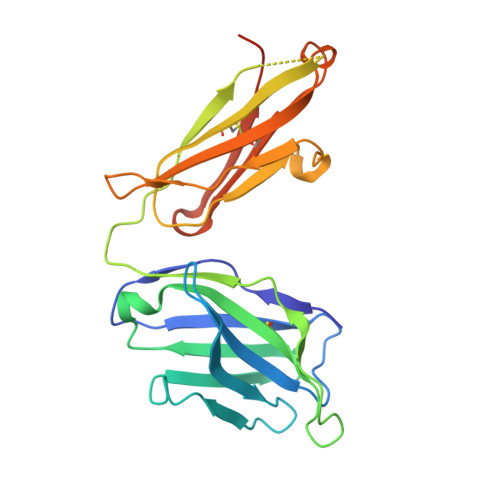
BAFF-neutralizing interaction of belimumab related to its therapeutic efficacy for treating systemic lupus erythematosus.
Shin, W., Lee, H.T., Lim, H., Lee, S.H., Son, J.Y., Lee, J.U., Yoo, K.Y., Ryu, S.E., Rhie, J., Lee, J.Y., Heo, Y.S.(2018) Nat Commun 9: 1200-1200
- PubMed: 29572471 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-03620-2
- Primary Citation Related Structures:
5Y9J, 5Y9K - PubMed Abstract:
BAFF, a member of the TNF superfamily, has been recognized as a good target for autoimmune diseases. Belimumab, an anti-BAFF monoclonal antibody, was approved by the FDA for use in treating systemic lupus erythematosus. However, the molecular basis of BAFF neutralization by belimumab remains unclear. Here our crystal structure of the BAFF-belimumab Fab complex shows the precise epitope and the BAFF-neutralizing mechanism of belimumab, and demonstrates that the therapeutic activity of belimumab involves not only antagonizing the BAFF-receptor interaction, but also disrupting the formation of the more active BAFF 60-mer to favor the induction of the less active BAFF trimer through interaction with the flap region of BAFF. In addition, the belimumab HCDR3 loop mimics the DxL(V/L) motif of BAFF receptors, thereby binding to BAFF in a similar manner as endogenous BAFF receptors. Our data thus provides insights for the design of new drugs targeting BAFF for the treatment of autoimmune diseases.
- Department of Chemistry, Konkuk University, 120 Neungdong-ro, Gwangjin-gu, Seoul, 05029, Republic of Korea.
Organizational Affiliation: