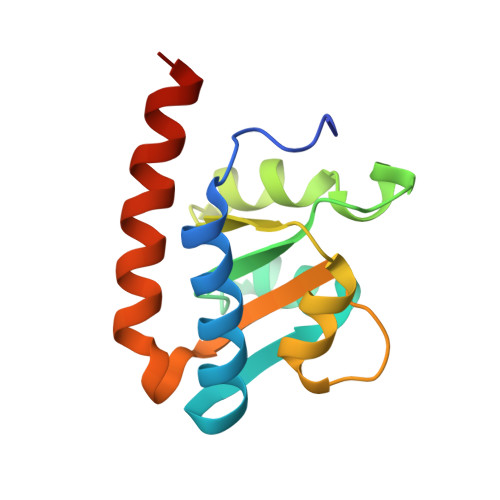
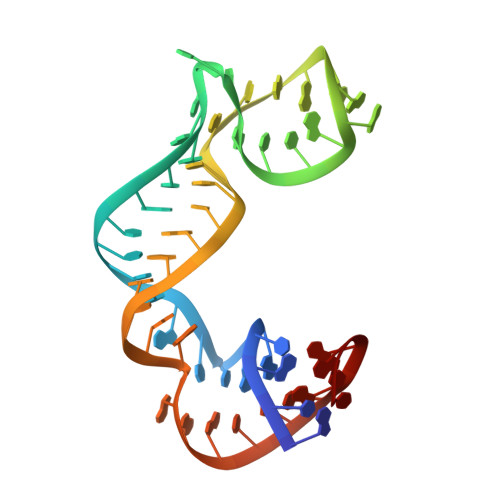
Crystal structures of the archaeal RNase P protein Rpp38 in complex with RNA fragments containing a K-turn motif.
Oshima, K., Gao, X., Hayashi, S., Ueda, T., Nakashima, T., Kimura, M.(2018) Acta Crystallogr F Struct Biol Commun 74: 57-64
- PubMed: 29372908 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X17018039
- Primary Citation Related Structures:
5XTM, 5Y7M - PubMed Abstract:
A characteristic feature of archaeal ribonuclease P (RNase P) RNAs is that they have extended helices P12.1 and P12.2 containing kink-turn (K-turn) motifs to which the archaeal RNase P protein Rpp38, a homologue of the human RNase P protein Rpp38, specifically binds. PhoRpp38 from the hyperthermophilic archaeon Pyrococcus horikoshii is involved in the elevation of the optimum temperature of the reconstituted RNase P by binding the K-turns in P12.1 and P12.2. Previously, the crystal structure of PhoRpp38 in complex with the K-turn in P12.2 was determined at 3.4 Å resolution. In this study, the crystal structure of PhoRpp38 in complex with the K-turn in P12.2 was improved to 2.1 Å resolution and the structure of PhoRpp38 in complex with the K-turn in P12.1 was also determined at a resolution of 3.1 Å. Both structures revealed that Lys35, Asn38 and Glu39 in PhoRpp38 interact with characteristic G·A and A·G pairs in the K-turn, while Thr37, Asp59, Lys84, Glu94, Ala96 and Ala98 in PhoRpp38 interact with the three-nucleotide bulge in the K-turn. Moreover, an extended stem-loop containing P10-P12.2 in complex with PhoRpp38, as well as PhoRpp21 and PhoRpp29, which are the archaeal homologues of the human proteins Rpp21 and Rpp29, respectively, was affinity-purified and crystallized. The crystals thus grown diffracted to a resolution of 6.35 Å. Structure determination of the crystals will demonstrate the previously proposed secondary structure of stem-loops including helices P12.1 and P12.2 and will also provide insight into the structural organization of the specificity domain in P. horikoshii RNase P RNA.
- Laboratory of Biochemistry, Department of Bioscience and Biotechnology, Graduate School, Faculty of Agriculture, Kyushu University, Hakozaki 6-10-1, Fukuoka 812-8581, Japan.
Organizational Affiliation: