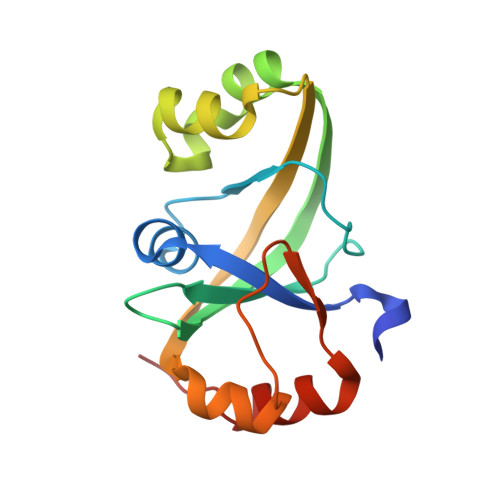
Crystal structure of an ASCH protein from Zymomonas mobilis and its ribonuclease activity specific for single-stranded RNA.
Kim, B.N., Shin, M., Ha, S.C., Park, S.Y., Seo, P.W., Hofmann, A., Kim, J.S.(2017) Sci Rep 7: 12303-12303
- PubMed: 28951575 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-12186-w
- Primary Citation Related Structures:
5GUQ, 5GUS, 5Y6B, 5Y6C - PubMed Abstract:
Activating signal cointegrator-1 homology (ASCH) domains were initially reported in human as a part of the ASC-1 transcriptional regulator, a component of a putative RNA-interacting protein complex; their presence has now been confirmed in a wide range of organisms. Here, we have determined the trigonal and monoclinic crystal structures of an ASCH domain-containing protein from Zymomonas mobilis (ZmASCH), and analyzed the structural determinants of its nucleic acid processing activity. The protein has a central β-barrel structure with several nearby α-helices. Positively charged surface patches form a cleft that runs through the pocket formed between the β-barrel and the surrounding α-helices. We further demonstrate by means of in vitro assays that ZmASCH binds nucleic acids, and degrades single-stranded RNAs in a magnesium ion-dependent manner with a cleavage preference for the phosphodiester bond between the pyrimidine and adenine nucleotides. ZmASCH also removes a nucleotide at the 5'-end. Mutagenesis studies, guided by molecular dynamics simulations, confirmed that three residues (Tyr47, Lys53, and Ser128) situated in the cleft contribute to nucleic acid-binding and RNA cleavage activities. These structural and biochemical studies imply that prokaryotic ASCH may function to control the cellular RNA amount.
- Department of Chemistry, Chonnam National University, Gwangju, 61186, Republic of Korea.
Organizational Affiliation: