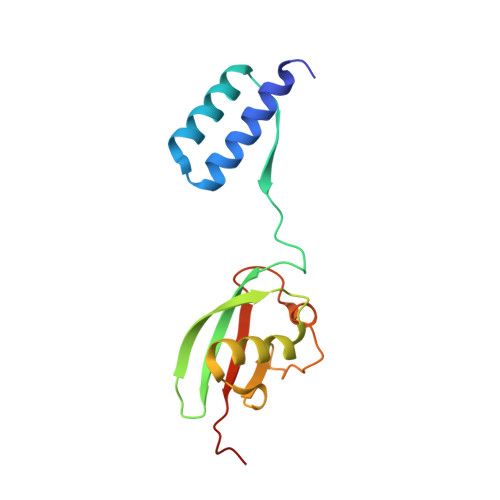
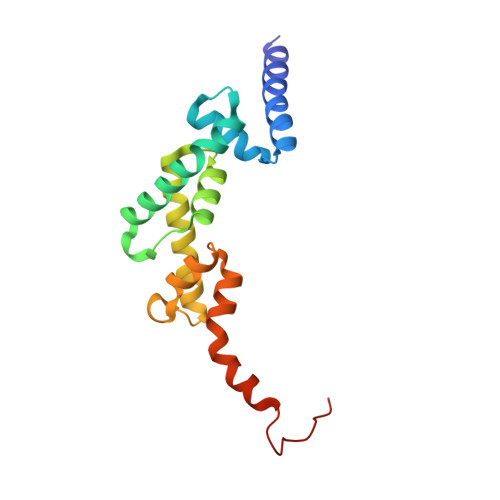
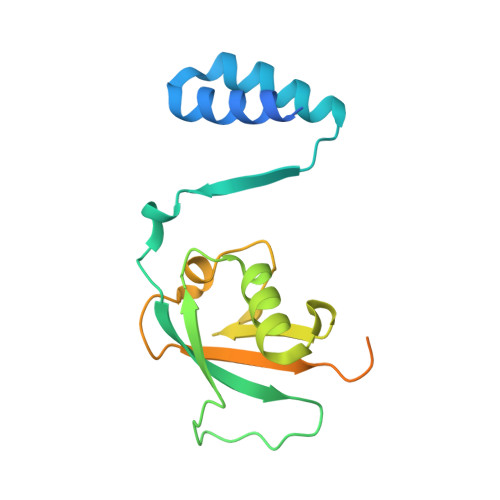
Cooperative Domain Formation by Homologous Motifs in HOIL-1L and SHARPIN Plays A Crucial Role in LUBAC Stabilization.
Fujita, H., Tokunaga, A., Shimizu, S., Whiting, A.L., Aguilar-Alonso, F., Takagi, K., Walinda, E., Sasaki, Y., Shimokawa, T., Mizushima, T., Ohki, I., Ariyoshi, M., Tochio, H., Bernal, F., Shirakawa, M., Iwai, K.(2018) Cell Rep 23: 1192-1204
- PubMed: 29694895 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2018.03.112
- Primary Citation Related Structures:
5Y3T - PubMed Abstract:
The linear ubiquitin chain assembly complex (LUBAC) participates in inflammatory and oncogenic signaling by conjugating linear ubiquitin chains to target proteins. LUBAC consists of the catalytic HOIP subunit and two accessory subunits, HOIL-1L and SHARPIN. Interactions between the ubiquitin-associated (UBA) domains of HOIP and the ubiquitin-like (UBL) domains of two accessory subunits are involved in LUBAC stabilization, but the precise molecular mechanisms underlying the formation of stable trimeric LUBAC remain elusive. We solved the co-crystal structure of the binding regions of the trimeric LUBAC complex and found that LUBAC-tethering motifs (LTMs) located N terminally to the UBL domains of HOIL-1L and SHARPIN heterodimerize and fold into a single globular domain. This interaction is resistant to dissociation and plays a critical role in stabilizing trimeric LUBAC. Inhibition of LTM-mediated HOIL-1L/SHARPIN dimerization profoundly attenuated the function of LUBAC, suggesting LTM as a superior target of LUBAC destabilization for anticancer therapeutics.
- Department of Molecular and Cellular Physiology, Kyoto University School of Medicine, Kyoto 606-8501, Japan.
Organizational Affiliation: