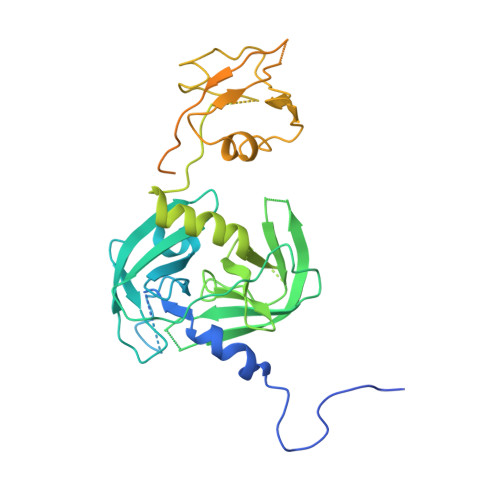
The unique trimeric assembly of the virulence factor HtrA fromHelicobacter pylorioccurs via N-terminal domain swapping.
Zhang, Z., Huang, Q., Tao, X., Song, G., Zheng, P., Li, H., Sun, H., Xia, W.(2019) J Biological Chem 294: 7990-8000
- PubMed: 30936204 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.007387
- Primary Citation Related Structures:
5Y28, 5Y2D - PubMed Abstract:
Knowledge of the molecular mechanisms of specific bacterial virulence factors can significantly contribute to antibacterial drug discovery. Helicobacter pylori is a Gram-negative microaerophilic bacterium that infects almost half of the world's population, leading to gastric disorders and even gastric cancer. H. pylori expresses a series of virulence factors in the host, among which high-temperature requirement A ( Hp HtrA) is a newly identified serine protease secreted by H. pylori. Hp HtrA cleaves the extracellular domain of the epithelial cell surface adhesion protein E-cadherin and disrupts gastric epithelial cell junctions, allowing H. pylori to access the intercellular space. Here we report the first crystal structure of Hp HtrA at 3.0 Å resolution. The structure revealed a new type of HtrA protease trimer stabilized by unique N-terminal domain swapping distinct from other known HtrA homologs. We further observed that truncation of the N terminus completely abrogates Hp HtrA trimer formation as well as protease activity. In the presence of unfolded substrate, Hp HtrA assembled into cage-like 12-mers or 24-mers. Combining crystallographic, biochemical, and mutagenic data, we propose a mechanistic model of how Hp HtrA recognizes and cleaves the well-folded E-cadherin substrate. Our study provides a fundamental basis for the development of anti- H. pylori agents by using a previously uncharacterized HtrA protease as a target.
- MOE Key Laboratory of Bioinorganic and Synthetic Chemistry, School of Chemistry, Sun Yat-sen University, Guangzhou, 510275, China.
Organizational Affiliation: