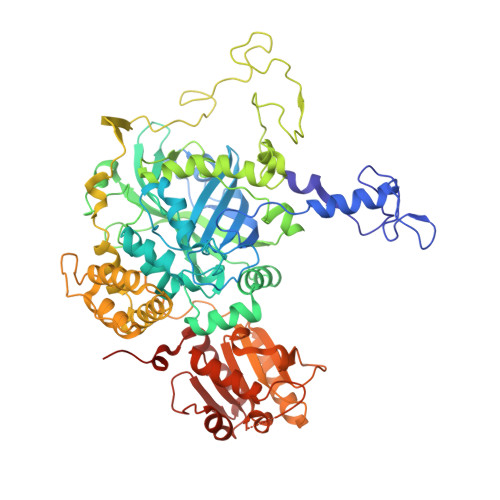
Identification of the site of oxidase substrate binding in Scytalidium thermophilum catalase.
Yuzugullu Karakus, Y., Goc, G., Balci, S., Yorke, B.A., Trinh, C.H., McPherson, M.J., Pearson, A.R.(2018) Acta Crystallogr D Struct Biol 74: 979-985
- PubMed: 30289408 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798318010628
- Primary Citation Related Structures:
5XVZ, 5XY4, 5Y17, 5ZZ1 - PubMed Abstract:
The catalase from Scytalidium thermophilum is a homotetramer containing a heme d in each active site. Although the enzyme has a classical monofunctional catalase fold, it also possesses oxidase activity towards a number of small organics, including catechol and phenol. In order to further investigate this, the crystal structure of the complex of the catalase with the classical catalase inhibitor 3-amino-1,2,4-triazole (3TR) was determined at 1.95 Å resolution. Surprisingly, no binding to the heme site was observed; instead, 3TR occupies a binding site corresponding to the NADPH-binding pocket in mammalian catalases at the entrance to a lateral channel leading to the heme. Kinetic analysis of site-directed mutants supports the assignment of this pocket as the binding site for oxidase substrates.
- Department of Biology, Kocaeli University, Umuttepe, 41380 Kocaeli, Turkey.
Organizational Affiliation: