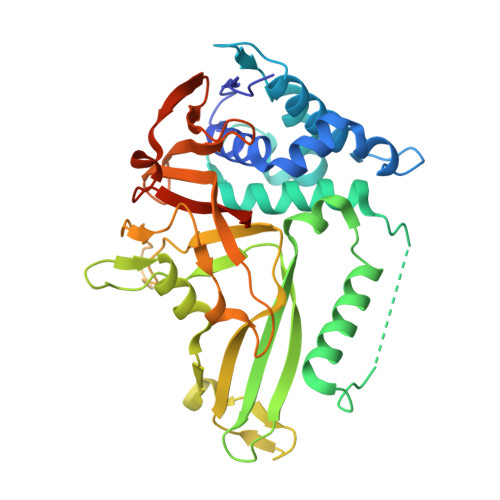
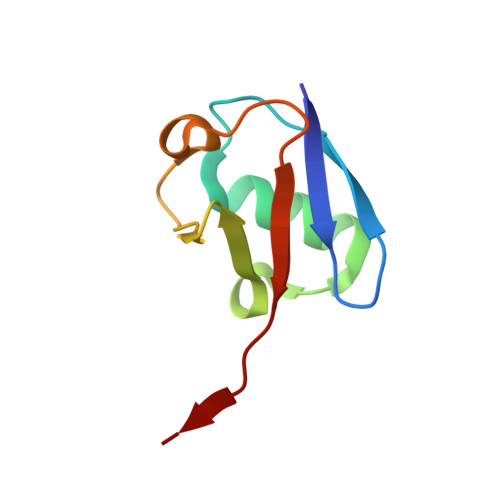
6-Thioguanine is a noncompetitive and slow binding inhibitor of human deubiquitinating protease USP2
Chuang, S.J., Cheng, S.C., Tang, H.C., Sun, C.Y., Chou, C.Y.(2018) Sci Rep 8: 3102-3102
- PubMed: 29449607 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-21476-w
- Primary Citation Related Structures:
5XU8, 5XVE - PubMed Abstract:
Ubiquitin-specific protease 2 (USP2) belongs to the family of deubiquitinases that can rescue protein targets from proteasomal degradation by reversing their ubiquitination. In various cancers, including prostate cancer and ovarian carcinoma, upregulation of USP2 leads to an increase in the levels of deubiquitinated substrates such as fatty acid synthase, MDM2, cyclin D1 and Aurora-A. USP2 thus plays a critical role in tumor cells' survival and therefore represents a therapeutic target. Here a leukemia drug, 6-thioguanine, was found to be a potent inhibitor of USP2. Enzyme-kinetic and X-ray crystallographic data suggest that 6-thioguanine displays a noncompetitive and slow-binding inhibitory mechanism against USP2. Our study provides a clear rationale for the clinical evaluation of 6-thioguanine for USP2-upregulated cancers.
- Department of Life Sciences and Institute of Genome Sciences, National Yang-Ming University, Taipei, 112, Taiwan.
Organizational Affiliation: