Discovery of Potent Vitamin D Receptor Antagonist
Kato, A., Tabei, M., Itoh, T., Yamamoto, K.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Vitamin D3 receptor | 271 | Rattus norvegicus | Mutation(s): 0 Gene Names: Vdr, Nr1i1 | 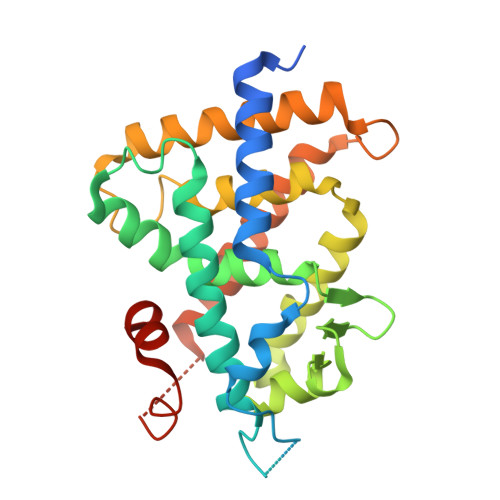 | |
UniProt | |||||
Find proteins for P13053 (Rattus norvegicus) Explore P13053 Go to UniProtKB: P13053 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P13053 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Find similar proteins by: Sequence | 3D Structure
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Mediator of RNA polymerase II transcription subunit 1 | B [auth C] | 13 | Homo sapiens | Mutation(s): 0 Gene Names: | 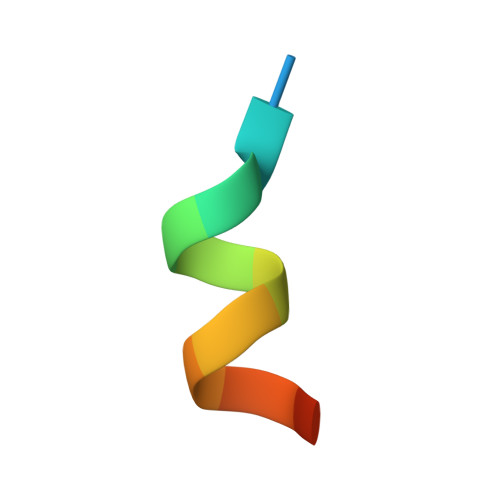 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for Q15648 (Homo sapiens) Explore Q15648 Go to UniProtKB: Q15648 | |||||
PHAROS: Q15648 GTEx: ENSG00000125686 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q15648 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 8FF Query on 8FF | C [auth A] | (4~{S})-4-[(1~{R})-1-[(1~{R},3~{a}~{S},4~{E},7~{a}~{R})-7~{a}-methyl-4-[2-[(3~{R},5~{R})-4-methylidene-3,5-bis(oxidanyl)cyclohexylidene]ethylidene]-2,3,3~{a},5,6,7-hexahydro-1~{H}-inden-1-yl]ethyl]-1-(1~{H}-pyrrol-2-yl)octan-1-one C33 H49 N O3 LSFVNCITLLLHBJ-LTRSXRPOSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 151.8 | α = 90 |
| b = 41.49 | β = 96.97 |
| c = 42.02 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| iMOSFLM | data reduction |
| Aimless | data scaling |
| REFMAC | phasing |