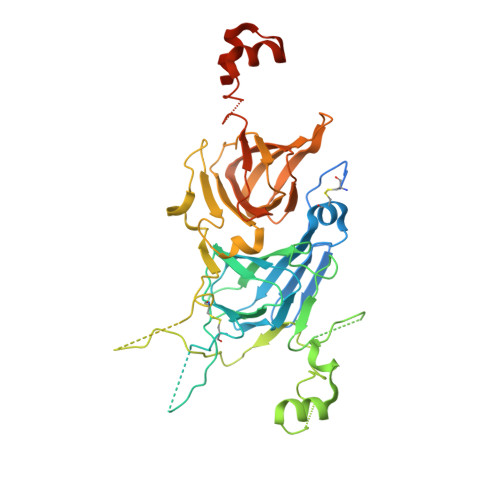
Crystal structure determination and analysis of 11S coconut allergen: Cocosin
Vajravijayan, S., Nandhagopal, N., Gunasekaran, K.(2017) Mol Immunol 92: 132-135
- PubMed: 29096167 Search on PubMed
- DOI: https://doi.org/10.1016/j.molimm.2017.10.018
- Primary Citation Related Structures:
5XTY - PubMed Abstract:
Allergy is an abnormal immune response against an innocuous target. Food allergy is an adverse reaction caused by common foods most well-known being those involving peanuts. Apart from mono sensitized food allergy, cross-reactivity with other food allergens is also commonly observed. To understand the phenomenon of cross-reactivity related to immune response, three dimensional structures of the allergens and their antigenic epitopes has to be analysed in detail. The X-ray crystal structure of Cocosin, a common 11S food allergen from coconut, has been determined at 2.2Å resolution using molecular replacement technique. The monomer of 52kDa is composed of two β-jelly roll domains, one with acidic and the other with basic character. The structure shows hexameric association with two trimers facing each other. Though the overall structure of Cocosin is similar to other 11S allergens, the occurrence of experimentally determined epitopes of the peanut allergen Ara h 3 at flexible as well as variable regions could be the reason for the clinically reported result of cross-reactivity that the peanut allergic patients are not sensitized with coconut allergen.
- Centre of Advanced Study in Crystallography and Biophysics, University of Madras, Guindy Campus, Chennai 600 025, India.
Organizational Affiliation: