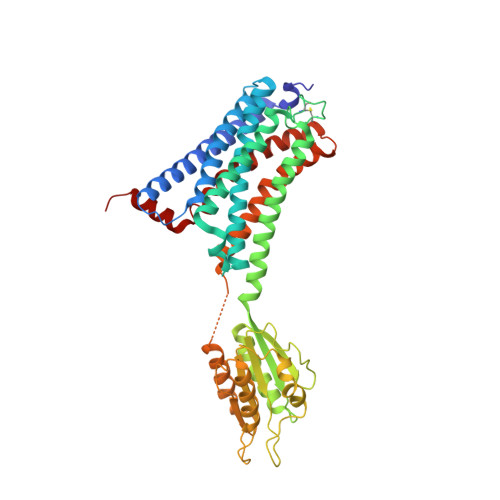
Crystal structures of agonist-bound human cannabinoid receptor CB1.
Hua, T., Vemuri, K., Nikas, S.P., Laprairie, R.B., Wu, Y., Qu, L., Pu, M., Korde, A., Jiang, S., Ho, J.H., Han, G.W., Ding, K., Li, X., Liu, H., Hanson, M.A., Zhao, S., Bohn, L.M., Makriyannis, A., Stevens, R.C., Liu, Z.J.(2017) Nature 547: 468-471
- PubMed: 28678776
- DOI: https://doi.org/10.1038/nature23272
- Primary Citation of Related Structures:
5XR8, 5XRA - PubMed Abstract:
The cannabinoid receptor 1 (CB 1 ) is the principal target of the psychoactive constituent of marijuana, the partial agonist Δ 9 -tetrahydrocannabinol (Δ 9 -THC). Here we report two agonist-bound crystal structures of human CB 1 in complex with a tetrahydrocannabinol (AM11542) and a hexahydrocannabinol (AM841) at 2.80 Å and 2.95 Å resolution, respectively. The two CB 1 -agonist complexes reveal important conformational changes in the overall structure, relative to the antagonist-bound state, including a 53% reduction in the volume of the ligand-binding pocket and an increase in the surface area of the G-protein-binding region. In addition, a 'twin toggle switch' of Phe200 3.36 and Trp356 6.48 (superscripts denote Ballesteros-Weinstein numbering) is experimentally observed and appears to be essential for receptor activation. The structures reveal important insights into the activation mechanism of CB 1 and provide a molecular basis for predicting the binding modes of Δ 9 -THC, and endogenous and synthetic cannabinoids. The plasticity of the binding pocket of CB 1 seems to be a common feature among certain class A G-protein-coupled receptors. These findings should inspire the design of chemically diverse ligands with distinct pharmacological properties.
- iHuman Institute, ShanghaiTech University, Shanghai 201210, China.
Organizational Affiliation: