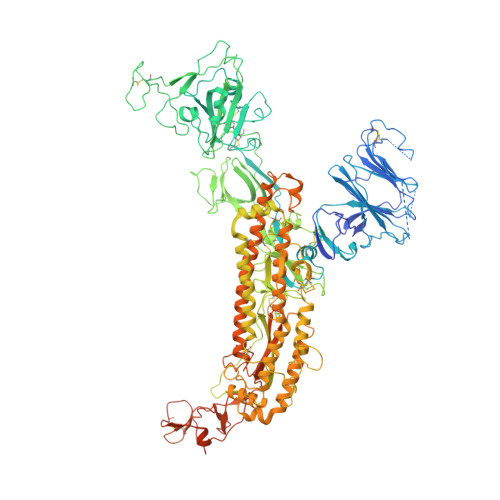
Cryo-electron microscopy structures of the SARS-CoV spike glycoprotein reveal a prerequisite conformational state for receptor binding.
Gui, M., Song, W., Zhou, H., Xu, J., Chen, S., Xiang, Y., Wang, X.(2017) Cell Res 27: 119-129
- PubMed: 28008928 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/cr.2016.152
- Primary Citation Related Structures:
5WRG, 5XLR - PubMed Abstract:
The global outbreak of SARS in 2002-2003 was caused by the infection of a new human coronavirus SARS-CoV. The infection of SARS-CoV is mediated mainly through the viral surface glycoproteins, which consist of S1 and S2 subunits and form trimer spikes on the envelope of the virions. Here we report the ectodomain structures of the SARS-CoV surface spike trimer in different conformational states determined by single-particle cryo-electron microscopy. The conformation 1 determined at 4.3 Å resolution is three-fold symmetric and has all the three receptor-binding C-terminal domain 1 (CTD1s) of the S1 subunits in "down" positions. The binding of the "down" CTD1s to the SARS-CoV receptor ACE2 is not possible due to steric clashes, suggesting that the conformation 1 represents a receptor-binding inactive state. Conformations 2-4 determined at 7.3, 5.7 and 6.8 Å resolutions are all asymmetric, in which one RBD rotates away from the "down" position by different angles to an "up" position. The "up" CTD1 exposes the receptor-binding site for ACE2 engagement, suggesting that the conformations 2-4 represent a receptor-binding active state. This conformational change is also required for the binding of SARS-CoV neutralizing antibodies targeting the CTD1. This phenomenon could be extended to other betacoronaviruses utilizing CTD1 of the S1 subunit for receptor binding, which provides new insights into the intermediate states of coronavirus pre-fusion spike trimer during infection.
- Center for Global Health and Infectious Diseases, Collaborative Innovation Center for Diagnosis and Treatment of Infectious Diseases, Beijing Advanced Innovation Center for Structural Biology, Department of Basic Medical Sciences, School of Medicine, Tsinghua University, Beijing 100084, China.
Organizational Affiliation: