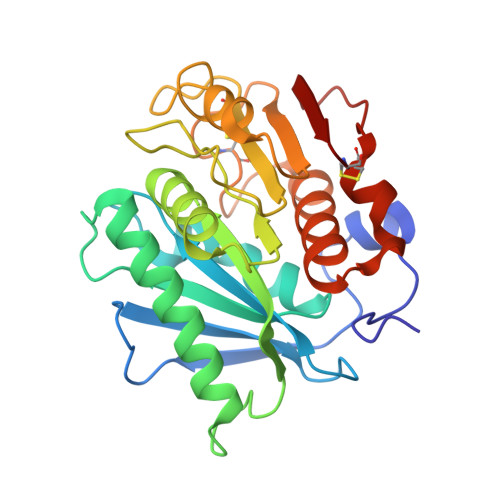
Structural insight into catalytic mechanism of PET hydrolase
Han, X., Liu, W., Huang, J.W., Ma, J., Zheng, Y., Ko, T.P., Xu, L., Cheng, Y.S., Chen, C.C., Guo, R.T.(2017) Nat Commun 8: 2106-2106
- PubMed: 29235460 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-02255-z
- Primary Citation Related Structures:
5XFY, 5XFZ, 5XG0, 5XH2, 5XH3 - PubMed Abstract:
PET hydrolase (PETase), which hydrolyzes polyethylene terephthalate (PET) into soluble building blocks, provides an attractive avenue for the bioconversion of plastics. Here we present the structures of a novel PETase from the PET-consuming microbe Ideonella sakaiensis in complex with substrate and product analogs. Through structural analyses, mutagenesis, and activity measurements, a substrate-binding mode is proposed, and several features critical for catalysis are elucidated.
- Industrial Enzymes National Engineering Laboratory, Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences, Tianjin, 300308, China.
Organizational Affiliation: