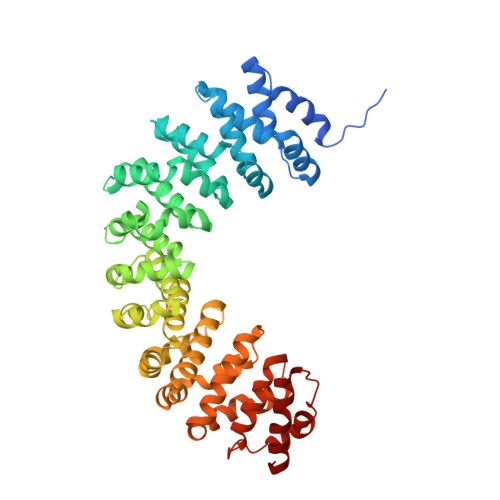
Structure-based analysis of the guanine nucleotide exchange factor SmgGDS reveals armadillo-repeat motifs and key regions for activity and GTPase binding
Shimizu, H., Toma-Fukai, S., Saijo, S., Shimizu, N., Kontani, K., Katada, T., Shimizu, T.(2017) J Biological Chem 292: 13441-13448
- PubMed: 28630045 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M117.792556
- Primary Citation Related Structures:
5XGC - PubMed Abstract:
Small GTPases are molecular switches that have critical biological roles and are controlled by GTPase-activating proteins and guanine nucleotide exchange factors (GEFs). The smg GDP dissociation stimulator (SmgGDS) protein functions as a GEF for the RhoA and RhoC small GTPases. SmgGDS has various regulatory roles, including small GTPase trafficking and localization and as a molecular chaperone, and interacts with many small GTPases possessing polybasic regions. Two SmgGDS splice variants, SmgGDS-558 and SmgGDS-607, differ in GEF activity and binding affinity for RhoA depending on the lipidation state, but the reasons for these differences are unclear. Here we determined the crystal structure of SmgGDS-558, revealing a fold containing tandem copies of armadillo repeats not present in other GEFs. We also observed that SmgGDS harbors distinct positively and negatively charged regions, both of which play critical roles in binding to RhoA and GEF activity. This is the first report demonstrating a relationship between the molecular function and atomic structure of SmgGDS. Our findings indicate that the two SmgGDS isoforms differ in GTPase binding and GEF activity, depending on the lipidation state, thus providing useful information about the cellular functions of SmgGDS in cells.
- From the Graduate School of Pharmaceutical Sciences, The University of Tokyo, 7-3-1 Hongo, Bunkyo-ku, Tokyo 113-0033, Japan.
Organizational Affiliation: