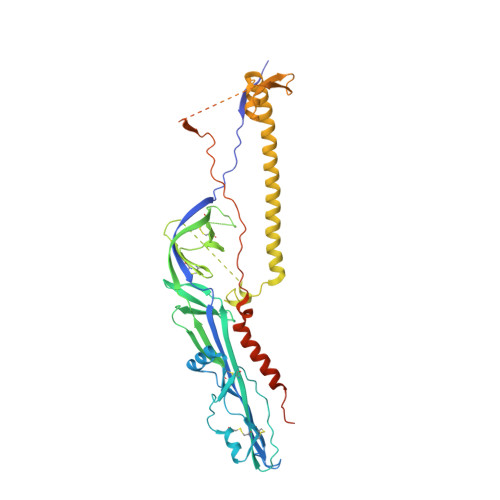
Structures of human-infecting Thogotovirus fusogens support a common ancestor with insect baculovirus
Peng, R., Zhang, S., Cui, Y., Shi, Y., Gao, G.F., Qi, J.(2017) Proc Natl Acad Sci U S A 114: E8905-E8912
- PubMed: 29073031 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1706125114
- Primary Citation Related Structures:
5XEA, 5XEB - PubMed Abstract:
Thogotoviruses are emerging tick-borne zoonotic orthomyxoviruses infecting both humans and domestic animals with severe clinical consequences. These viruses utilize a single-envelope glycoprotein (Gp) to facilitate their entry into host cells. Here, we present the Gp structures of Thogoto and Dhori viruses, both of which are members of the Thogotovirus genus in the family Orthomyxoviridae These structures, determined in the postfusion conformation, identified them as class III viral fusion proteins. It is intriguing that the Gp structures are similar to the envelope protein of baculovirus, although sharing a low sequence identity of ∼28%. Detailed structural and phylogenic analyses demonstrated that these Gps originated from a common ancestor. Among the structures, domain I is the most conserved region, particularly the fusion loops. Domain II showed the highest variability among different viruses, which might be related to their distinct host tropism. These findings increase our understanding of the divergent evolution processes of various orthomyxoviruses and indicate potential targets for developing antiviral therapeutics by intercepting virus entry.
- Chinese Academy of Sciences Key Laboratory of Pathogenic Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences, Beijing 100101, China.
Organizational Affiliation: