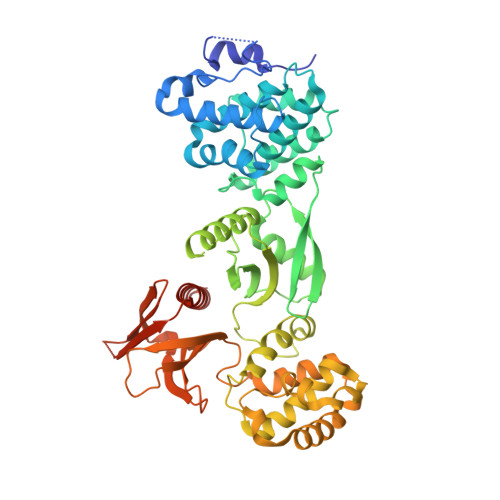
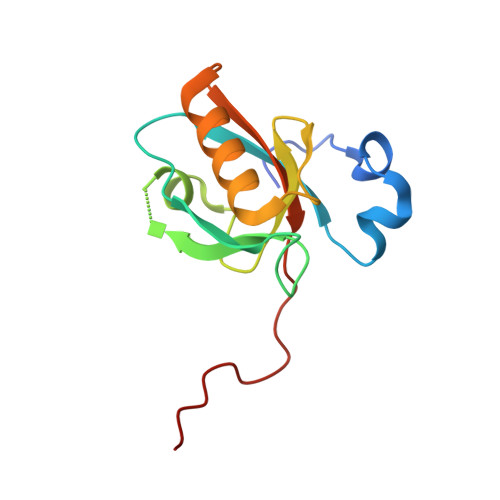
Structure of Myo7b/USH1C complex suggests a general PDZ domain binding mode by MyTH4-FERM myosins.
Li, J., He, Y., Weck, M.L., Lu, Q., Tyska, M.J., Zhang, M.(2017) Proc Natl Acad Sci U S A 114: E3776-E3785
- PubMed: 28439001 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1702251114
- Primary Citation Related Structures:
5XBF - PubMed Abstract:
Unconventional myosin 7a (Myo7a), myosin 7b (Myo7b), and myosin 15a (Myo15a) all contain MyTH4-FERM domains (myosin tail homology 4-band 4.1, ezrin, radixin, moesin; MF) in their cargo binding tails and are essential for the growth and function of microvilli and stereocilia. Numerous mutations have been identified in the MyTH4-FERM tandems of these myosins in patients suffering visual and hearing impairment. Although a number of MF domain binding partners have been identified, the molecular basis of interactions with the C-terminal MF domain (CMF) of these myosins remains poorly understood. Here we report the high-resolution crystal structure of Myo7b CMF in complex with the extended PDZ3 domain of USH1C (a.k.a., Harmonin), revealing a previously uncharacterized interaction mode both for MyTH4-FERM tandems and for PDZ domains. We predicted, based on the structure of the Myo7b CMF/USH1C PDZ3 complex, and verified that Myo7a CMF also binds to USH1C PDZ3 using a similar mode. The structure of the Myo7b CMF/USH1C PDZ complex provides mechanistic explanations for >20 deafness-causing mutations in Myo7a CMF. Taken together, these findings suggest that binding to PDZ domains, such as those from USH1C, PDZD7, and Whirlin, is a common property of CMFs of Myo7a, Myo7b, and Myo15a.
- Division of Life Science, State Key Laboratory of Molecular Neuroscience, Hong Kong University of Science and Technology, Clear Water Bay, Kowloon, Hong Kong.
Organizational Affiliation: